Calculate Effect Sizes and Outcome Measures
escalc.RdFunction to calculate various effect sizes or outcome measures (and the corresponding sampling variances) that are commonly used in meta-analyses.
escalc(measure, ai, bi, ci, di, n1i, n2i, x1i, x2i, t1i, t2i,
m1i, m2i, sd1i, sd2i, xi, mi, ri, ti, fi, pi,
sdi, r2i, ni, yi, vi, sei,
data, slab, flip, subset, include,
add=1/2, to="only0", drop00=FALSE,
vtype="LS", correct=TRUE, cutoff,
var.names=c("yi","vi"), add.measure=FALSE,
append=TRUE, replace=TRUE, digits, ...)Arguments
- measure
a character string to specify which effect size or outcome measure should be calculated (e.g.,
"SMD","ZCOR","OR"). See ‘Details’ for possible options and how the data needed to compute the selected effect size or outcome measure should then be specified (i.e., which of the following arguments need to be used).
These arguments pertain to data input:
- ai
vector with the \(2 \times 2\) table frequencies (upper left cell).
- bi
vector with the \(2 \times 2\) table frequencies (upper right cell).
- ci
vector with the \(2 \times 2\) table frequencies (lower left cell).
- di
vector with the \(2 \times 2\) table frequencies (lower right cell).
- n1i
vector with the group sizes or row totals (first group/row).
- n2i
vector with the group sizes or row totals (second group/row).
- x1i
vector with the number of events (first group).
- x2i
vector with the number of events (second group).
- t1i
vector with the total person-times (first group).
- t2i
vector with the total person-times (second group).
- m1i
vector with the means (first group or time point).
- m2i
vector with the means (second group or time point).
- sd1i
vector with the standard deviations (first group or time point).
- sd2i
vector with the standard deviations (second group or time point).
- xi
vector with the frequencies of the event of interest.
- mi
vector with the frequencies of the complement of the event of interest or the group means.
- ri
vector with the raw correlation coefficients.
- ti
vector with the total person-times or t-test statistics.
- fi
vector with the F-test statistics.
- pi
vector with the (signed) p-values.
- sdi
vector with the standard deviations.
- r2i
vector with the \(R^2\) values.
- ni
vector with the sample/group sizes.
- yi
vector with the effect size estimates.
- vi
vector with the corresponding sampling variances.
- sei
vector with the corresponding standard errors.
- data
optional data frame containing the variables given to the arguments above.
- slab
optional vector with labels for the studies.
- flip
optional logical to indicate whether to flip the sign of the estimates. Can also be a vector. Can also be a numeric vector to specify a multiplier.
- subset
optional (logical or numeric) vector to specify the subset of studies that will be included in the data frame returned by the function.
- include
optional (logical or numeric) vector to specify the subset of studies for which the measure should be calculated. See the ‘Value’ section for more details.
These arguments pertain to handling of zero cells/counts/frequencies:
- add
a non-negative number to specify the amount to add to zero cells, counts, or frequencies. See ‘Details’ and ‘Note’.
- to
a character string to specify when the values under
addshould be added (either"all","only0","if0all", or"none"). See ‘Details’.- drop00
logical to specify whether studies with no cases/events (or only cases) in both groups should be dropped when calculating the estimates. See ‘Details’.
These arguments pertain to the computations:
- vtype
a character string to specify the type of sampling variances to calculate. Can also be a vector. See ‘Details’.
- correct
logical to specify whether a bias correction should be applied to the estimates (the default is
TRUE).- cutoff
optional numeric value to adjust the cutoff for flagging large estimates. See ‘Details’.
These arguments pertain to the formatting of the returned data frame:
- var.names
character vector with two elements to specify the name of the variable for the estimates and the name of the variable for the corresponding sampling variances (the defaults are
"yi"and"vi").- add.measure
logical to specify whether a variable should be added to the data frame (with default name
"measure") that indicates the type of measure computed. When using this option,var.namescan have a third element to change this variable name.- append
logical to specify whether the data frame provided via the
dataargument should be returned together with the estimates and corresponding sampling variances (the default isTRUE).- replace
logical to specify whether existing values for
yiandviin the data frame should be replaced. Only relevant whenappend=TRUEand the data frame already contains theyiandvivariables. Ifreplace=TRUE(the default), all of the existing values will be overwritten. Ifreplace=FALSE, onlyNAvalues will be replaced. See the ‘Value’ section for more details.- digits
optional integer to specify the number of decimal places to which the printed results should be rounded. If unspecified, the default is 4. Note that the values are stored without rounding in the returned object. See also here for further details on how to control the number of digits in the output.
- ...
other arguments.
Details
Before a meta-analysis can be conducted, the relevant results from each study must be quantified in such a way that the resulting values can be further aggregated and compared. Depending on (a) the goals of the meta-analysis, (b) the design and types of studies included, and (c) the information provided therein, one of the various effect sizes or ‘outcome measures’ described below may be appropriate for the meta-analysis and can be computed with the escalc function.
The measure argument is a character string to specify the measure that should be calculated (see below for the various options), arguments ai through ni are then used to specify the information needed to calculate the various measures (depending on the chosen measure, different arguments need to be specified), and data can be used to specify a data frame containing the variables given to the previous arguments. The add, to, and drop00 arguments may be needed when dealing with frequency or count data that needs special handling when some of the frequencies or counts are equal to zero (see below for details). Finally, the vtype argument is used to specify how the sampling variances should be computed (again, see below for details).
To provide a structure to the measures that can be calculated with the escalc function, we can distinguish between measures that are used to:
| (1) | contrast two independent (either experimentally created or naturally occurring) groups, | |
| (2) | describe the direction and strength of the association between two variables, | |
| (3) | summarize some characteristic or attribute of individual groups, or | |
| (4) | quantify change within a single group or the difference between two matched/paired samples. |
Furthermore, where appropriate, we can further distinguish between measures that are applicable when the characteristic, response, or dependent variable assessed within the individual studies is:
| (a) | a quantitative variable (e.g., amount of depression as assessed by a rating scale), | |
| (b) | a dichotomous (binary) variable (e.g., remission versus no remission), | |
| (c) | a count of events per time unit (e.g., number of migraines per year), or | |
| (d) | a mix of the types above. |
Below, these number and letter codes are used (also in combination) to make it easier to quickly find a measure suitable for a particular meta-analysis (e.g., search for (1b) to find measures that describe the difference between two groups with respect to a dichotomous variable or (2a) for measures that quantify the association between two quantitative variables).
(1) Measures for Two-Group Comparisons
In many meta-analyses, the goal is to synthesize the results from studies that compare or contrast two groups. The groups may be experimentally defined (e.g., a treatment and a control group created via random assignment) or may occur naturally (e.g., men and women, employees working under high- versus low-stress conditions, people/animals/plants exposed to some environmental risk factor versus those not exposed, patients versus controls).
(1a) Measures for Quantitative Variables
When the response or dependent variable assessed within the individual studies is measured on a quantitative scale, it is customary to report certain summary statistics, such as the mean and standard deviation of the observations within the two groups (in case medians, min/max values, and quartiles are reported, see conv.fivenum for a function that can be used to estimate means and standard deviations from such statistics). The data layout for a study comparing two groups with respect to such a variable is then of the form:
| mean | standard deviation | group size | ||||
| group 1 | m1i | sd1i | n1i | |||
| group 2 | m2i | sd2i | n2i |
where m1i and m2i are the observed means of the two groups, sd1i and sd2i are the observed standard deviations, and n1i and n2i denote the number of individuals in each group.
Measures for Differences in Central Tendency
Often, interest is focused on differences between the two groups with respect to their central tendency. The raw mean difference, the standardized mean difference, and the (log transformed) ratio of means (also called the log ‘response ratio’) are useful measures when meta-analyzing studies of this type.
The options for the measure argument are then:
"MD"for the raw mean difference (e.g., Borenstein, 2009),"SMD"for the standardized mean difference (Hedges, 1981),"SMDH"for the standardized mean difference with heteroscedastic population variances in the two groups (Bonett, 2008, 2009),"SMD1"for the standardized mean difference where the mean difference is divided by the standard deviation of the second group (and"SMD1H"for the same but with heteroscedastic population variances),"ROM"for the log transformed ratio of means (Hedges et al., 1999; Lajeunesse, 2011).
The raw mean difference is simply \((\text{m1i}-\text{m2i})\), while the standardized mean difference is given by \((\text{m1i}-\text{m2i})/\text{sdi}\). For measure="SMD", \(\text{sdi} = \sqrt{\frac{(\text{n1i}-1)\text{sd1i}^2 + (\text{n2i}-1)\text{sd2i}^2}{\text{n1i}+\text{n2i}-2}}\) is the pooled standard deviation of the two groups (assuming homoscedasticity of the population variances). For measure="SMDH", \(\text{sdi} = \sqrt{\frac{\text{sd1i}^2 + \text{sd2i}^2}{2}}\) is the square root of the average variance (allowing for heteroscedastic population variances). Finally, for measure="SMD1" and measure="SMD1H", \(\text{sdi} = \text{sd2i}\) (note: for measure="SMD1", only sd2i needs to be specified and sd1i is ignored).
For measure="SMD", the positive bias in the standardized mean difference (i.e., in a Cohen's d value) is automatically corrected for within the function, yielding Hedges' g (Hedges, 1981). Similarly, the analogous bias correction is applied for measure="SMDH" (Bonett, 2009), measure="SMD1" (Hedges, 1981), and measure="SMD1H". With correct=FALSE, these bias corrections can be switched off.
For measure="ROM", the log is taken of the ratio of means (i.e., \(\log(\text{m1i}/\text{m2i})\)), which makes this measure symmetric around 0 and results in a sampling distribution that is closer to normality. Hence, this measure cannot be computed when m1i and m2i have opposite signs (in fact, this measure is only meant to be used for ratio scale measurements, where both means should be positive anyway). A bias correction is also applied to this measure (Lajeunesse, 2015, equation 8) unless correct=FALSE.
For measure="SMD", if the means and standard deviations are unknown for some studies, various other inputs can be used to recover the standardized mean differences. In the case that the standardized mean differences (Cohen's d values) are directly available (e.g., they are reported in some studies), then these can be specified via argument di. If the point-biserial correlations (between the group dummy variable and the quantitative response/dependent variable) are known, these can be specified via argument ri. If the t-statistics from an independent samples (Student's) t-test are available, these can be specified via argument ti. Note that the sign of these inputs is then taken to be the sign of the standardized mean differences. If only the two-sided p-values corresponding to the t-tests are known, one can specify those values via argument pi (which are then transformed into the t-statistics and then further into the standardized mean differences). However, since a two-sided p-value does not carry information about the sign of the test statistic (and hence neither about the standardized mean difference), the sign of the p-values (which can be negative) is used as the sign of the standardized mean differences (e.g., escalc(measure="SMD", pi=-0.018, n1i=20, n2i=20) yields a negative standardized mean difference of -0.7664). See here for a more detailed illustration of using the di, ti, and pi arguments.
For measure="MD", one can choose between vtype="LS" (the default) and vtype="HO". The former computes the sampling variances without assuming homoscedasticity (i.e., that the true variances of the measurements are the same in group 1 and group 2 within each study), while the latter assumes homoscedasticity (equations 12.5 and 12.3 in Borenstein, 2009, respectively). For measure="SMD", one can choose between vtype="LS" (the default) for the usual large-sample approximation to compute the sampling variances (equation 8 in Hedges, 1982), vtype="LS2" to compute the sampling variances as described in Borenstein (2009; equation 12.17), vtype="UB" to compute unbiased estimates of the sampling variances (equation 9 in Hedges, 1983), vtype="AV" to compute the sampling variances with the usual large-sample approximation but plugging the sample-size weighted average of the Hedges' g values into the equation, and vtype="H0" to compute the sampling variances under the null hypothesis (that the true standardized mean differences are equal to zero). The same choices also apply to measure="SMD1". For measure="ROM", one can choose between vtype="LS" (the default) for the usual large-sample approximation to compute the sampling variances (equation 1 in Hedges et al., 1999), vtype="HO" to compute the sampling variances assuming homoscedasticity (the unnumbered equation after equation 1 in Hedges et al., 1999), vtype="LS2" to compute the sampling variances based on the second-order Taylor expansion (equation 9 in Lajeunesse, 2015), vtype="AV" to compute the sampling variances assuming homoscedasticity of the coefficient of variation within each group across studies, vtype="AVHO" to compute the sampling variances assuming homoscedasticity of the coefficient of variation for both groups across studies (see Nakagawa et al., 2023, for details on the latter two options and why they can be advantageous).
Datasets corresponding to data of this type are provided in dat.normand1999, dat.curtis1998, and dat.gibson2002.
Measures for Variability Differences
Interest may also be focused on differences between the two groups with respect to their variability. For this, the (log transformed) ratio of the standard deviations (also called the ‘variability ratio’) can be used (Nakagawa et al., 2015). To also account for differences in mean levels, the (log transformed) ratio between the coefficients of variation from the two groups (also called the ‘coefficient of variation ratio’) can be a useful measure (Nakagawa et al., 2015). The options for the measure argument are:
"VR"for the log transformed variability ratio,"CVR"for the log transformed coefficient of variation ratio.
Measure "VR" is computed with \(\log(\text{sd1i}/\text{sd2i})\) and hence one only needs to specify sd1i, sd2i, n1i, and n2i (i.e., arguments m1i and m2i are irrelevant). Measure "CVR" is computed with \(\log\mathopen{}\left(\left(\text{sd1i}/\text{m1i}\right) \middle/ \left(\text{sd2i}/\text{m2i}\right) \right)\mathclose{}\). Note that a bias correction is applied for both of these measures (Senior et al., 2020, equations 5 and 6) unless correct=FALSE. When vtype="LS" (the default), then the sampling variances are computed with equations 11 and 13 from Senior et al. (2020) for "VR" and "CVR", respectively. When vtype="LS2", then equations 15 and 16 (with a slight correction, multiplying the third and sixth term by 1/2) based on the second-order Taylor expansions are used instead.
Measures for Stochastic Superiority
Another way to quantify the difference between two groups is in terms of the ‘common language effect size’ (CLES) (McGraw & Wong, 1992). This measure provides an estimate of \(P(X \gt Y)\), that is, the probability that a randomly chosen person from the first group has a larger value on the response variable than a randomly chosen person from the second group (or in case \(X\) and \(Y\) values can be tied, we define the measure as \(P(X \gt Y) + \frac{1}{2} P(X = Y)\)). This measure is identical to the area under the curve (AUC) under the receiver operating characteristic (ROC) curve (e.g., for a diagnostic test or more broadly for a binary classifier) and the ‘concordance probability’ (or c-statistic) and is directly related to the \(U\) statistic from the Mann-Whitney U test (i.e., \(\text{CLES} = U / (n_1 \times n_2)\)).
If the CLES/AUC values with corresponding sampling variances (or standard errors) are known, they can be directly meta-analyzed for example using the rma.uni function. However, in practice, one is likely to encounter studies that only report CLES/AUC values and the group sizes. In this case, one can specify these values via the ai, n1i, and n2i arguments and set measure="CLES" (or equivalently measure="AUC"). If vtype="LS" (the default), the sampling variances are then computed based on Newcombe (2006) (method 4), but using (n1i-1)(n2i-1) in the denominator as suggested by Cho et al. (2019). If vtype="LS2", the sampling variances are computed based on Hanley and McNeil (1982; equations 1 and 2), again using (n1i-1)(n2i-1) in the denominator (and in the unlikely case that the proportion of tied values is known, this can be specified via argument mi, in which case the adjustment as described by Cho et al. (2019) is also applied).
Under the assumption that the data within the two groups are normally distributed (the so-called binormal model), one can also estimate the CLES/AUC values from the means and standard deviations of the two groups. For this, one sets measure="CLESN" (or equivalently measure="AUCN") and specifies the means via arguments m1i and m2i, the standard deviations via arguments sd1i and sd2i, and the group sizes via arguments n1i and n2i. If vtype="LS" (the default), the sampling variances are then computed based on the large-sample approximation derived via the delta method (equation 3 (with a correction, since the plus sign in front of the last term in braces should be a multiplication sign) and equation 4 in Goddard & Hinberg, 1990, but using n1i-1 and n2i-1 in the denominators). Computing the CLES/AUC values and corresponding sampling variances does not assume homoscedasticity of the variances in the two groups. However, when vtype="HO", then homoscedasticity is assumed (this will also affect the calculation of the CLES/AUC values themselves). As for measure="SMD", one can also specify standardized mean differences via argument di, t-statistics from an independent samples t-test via argument ti, and/or signed two-sided p-values corresponding to the t-tests via argument pi, which all can be converted into CLES/AUC values (note that this automatically assumes homoscedasticity). One can also directly specify binormal model CLES/AUC values via argument ai (but unless the corresponding sd1i and sd2i values are also specified, the sampling variances are then computed under the assumption of homoscedasticity).
(1b) Measures for Dichotomous Variables
In various fields of research (such as the health and medical sciences), the response variable measured within the individual studies is often dichotomous (binary), so that the data from a study comparing two different groups can be expressed in terms of a \(2 \times 2\) table, such as:
| outcome 1 | outcome 2 | total | ||||
| group 1 | ai | bi | n1i | |||
| group 2 | ci | di | n2i |
where ai, bi, ci, and di denote the cell frequencies (i.e., the number of individuals falling into a particular category) and n1i and n2i are the row totals (i.e., the group sizes).
For example, in a set of randomized clinical trials, group 1 and group 2 may refer to the treatment and placebo/control group, respectively, with outcome 1 denoting some event of interest (e.g., death, complications, failure to improve under the treatment) and outcome 2 its complement. Similarly, in a set of cohort studies, group 1 and group 2 may denote those who engage in and those who do not engage in a potentially harmful behavior (e.g., smoking), with outcome 1 denoting the development of a particular disease (e.g., lung cancer) during the follow-up period. Finally, in a set of case-control studies, group 1 and group 2 may refer to those with the disease (i.e., cases) and those free of the disease (i.e., controls), with outcome 1 denoting, for example, exposure to some environmental risk factor in the past and outcome 2 non-exposure. Note that in all of these examples, the stratified sampling scheme fixes the row totals (i.e., the group sizes) by design.
A meta-analysis of studies reporting results in terms of \(2 \times 2\) tables can be based on one of several different measures, including the risk ratio (also called the relative risk), the odds ratio, the risk difference, and the arcsine square root transformed risk difference (e.g., Fleiss & Berlin, 2009, Rücker et al., 2009). For any of these measures, one needs to specify the cell frequencies via the ai, bi, ci, and di arguments (or alternatively, one can use the ai, ci, n1i, and n2i arguments).
The options for the measure argument are then:
"RR"for the log risk ratio,"OR"for the log odds ratio,"RD"for the risk difference,"AS"for the arcsine square root transformed risk difference (Rücker et al., 2009),"PETO"for the log odds ratio estimated with Peto's method (Yusuf et al., 1985).
Let \(\text{p1i} = \text{ai}/\text{n1i}\) and \(\text{p2i} = \text{ci}/\text{n2i}\) denote the proportion of individuals with outcome 1 in group 1 and group 2, respectively. Then the log risk ratio is computed with \(\log(\text{p1i}/\text{p2i})\), the log odds ratio with \(\log\mathopen{}\left(\left(\frac{\text{p1i}}{1-\text{p1i}}\right) \middle/ \left(\frac{\text{p2i}}{1-\text{p2i}}\right) \right)\mathclose{}\), the risk difference with \(\text{p1i}-\text{p2i}\), and the arcsine square root transformed risk difference with \(\text{asin}(\sqrt{\text{p1i}})-\text{asin}(\sqrt{\text{p2i}})\). See Yusuf et al. (1985) for the computation of the log odds ratio when measure="PETO". Note that the log is taken of the risk ratio and the odds ratio, which makes these measures symmetric around 0 and results in corresponding sampling distributions that are closer to normality. Also, when multiplied by 2, the arcsine square root transformed risk difference is identical to Cohen's h (Cohen, 1988). For all of these measures, a positive value indicates that the proportion of individuals with outcome 1 is larger in group 1 compared to group 2.
Cell entries with a zero count can be problematic, especially for the risk ratio and the odds ratio. Adding a small constant to the cells of the \(2 \times 2\) tables is a common solution to this problem. When to="only0" (the default), the value of add (the default is 1/2; but see ‘Note’) is added to each cell of those \(2 \times 2\) tables with at least one cell equal to 0. When to="all", the value of add is added to each cell of all \(2 \times 2\) tables. When to="if0all", the value of add is added to each cell of all \(2 \times 2\) tables, but only when there is at least one \(2 \times 2\) table with a zero cell. Setting to="none" or add=0 has the same effect: No adjustment to the observed table frequencies is made. Depending on the measure and the data, this may lead to division by zero (when this occurs, the resulting value is recoded to NA). Also, studies where ai=ci=0 or bi=di=0 may be considered to be uninformative about the size of the effect and dropping such studies has sometimes been recommended (Higgins et al., 2019). This can be done by setting drop00=TRUE. The values for such studies will then be set to NA (i.e., missing).
Datasets corresponding to data of this type are provided in dat.bcg, dat.collins1985a, dat.collins1985b, dat.egger2001, dat.hine1989, dat.laopaiboon2015, dat.lee2004, dat.li2007, dat.linde2005, dat.nielweise2007, and dat.yusuf1985.
If the \(2 \times 2\) table is not available (or cannot be reconstructed, for example with the conv.2x2 function) for a study, but the odds ratio and the corresponding confidence interval is reported, one can easily transform these values into the corresponding log odds ratio and sampling variance (and combine such a study with those that do report \(2 \times 2\) table data). See the conv.wald function and here for an illustration/discussion of this.
(1c) Measures for Event Counts
In medical and epidemiological studies comparing two different groups (e.g., treated versus untreated patients, exposed versus unexposed individuals), results are sometimes reported in terms of event counts (i.e., the number of events, such as strokes or myocardial infarctions) over a certain period of time. Data of this type are also referred to as ‘person-time data’. Assume that the studies report data in the form:
| number of events | total person-time | |||
| group 1 | x1i | t1i | ||
| group 2 | x2i | t2i |
where x1i and x2i denote the number of events in the first and the second group, respectively, and t1i and t2i the corresponding total person-times at risk. Often, the person-time is measured in years, so that t1i and t2i denote the total number of follow-up years in the two groups.
This form of data is fundamentally different from what was described in the previous section, since the total follow-up time may differ even for groups of the same size and the individuals studied may experience the event of interest multiple times. Hence, different measures than the ones described in the previous section need to be considered when data are reported in this format. These include the incidence rate ratio, the incidence rate difference, and the square root transformed incidence rate difference (Bagos & Nikolopoulos, 2009; Rothman et al., 2008). For any of these measures, one needs to specify the total number of events via the x1i and x2i arguments and the corresponding total person-time values via the t1i and t2i arguments.
The options for the measure argument are then:
"IRR"for the log incidence rate ratio,"IRD"for the incidence rate difference,"IRSD"for the square root transformed incidence rate difference.
Let \(\text{ir1i} = \text{x1i}/\text{t1i}\) and \(\text{ir2i} = \text{x2i}/\text{t2i}\) denote the observed incidence rates in each group. Then the log incidence rate ratio is computed with \(\log(\text{ir1i}/\text{ir2i})\), the incidence rate difference with \(\text{ir1i}-\text{ir2i}\), and the square root transformed incidence rate difference with \(\sqrt{\text{ir1i}}-\sqrt{\text{ir2i}}\). Note that the log is taken of the incidence rate ratio, which makes this measure symmetric around 0 and results in a sampling distribution that is closer to normality.
Studies with zero events in one or both groups can be problematic, especially for the incidence rate ratio. Adding a small constant to the number of events is a common solution to this problem. When to="only0" (the default), the value of add (the default is 1/2; but see ‘Note’) is added to x1i and x2i only in the studies that have zero events in one or both groups. When to="all", the value of add is added to x1i and x2i in all studies. When to="if0all", the value of add is added to x1i and x2i in all studies, but only when there is at least one study with zero events in one or both groups. Setting to="none" or add=0 has the same effect: No adjustment to the observed number of events is made. Depending on the measure and the data, this may lead to division by zero (when this occurs, the resulting value is recoded to NA). Like for \(2 \times 2\) table data, studies where x1i=x2i=0 may be considered to be uninformative about the size of the effect and dropping such studies has sometimes been recommended. This can be done by setting drop00=TRUE. The values for such studies will then be set to NA.
Datasets corresponding to data of this type are provided in dat.hart1999 and dat.nielweise2008.
(1d) Transforming SMDs to ORs and Vice-Versa
In some meta-analyses, one may encounter studies that contrast two groups with respect to a quantitative response variable (case 1a above) and other studies that contrast the same two groups with respect to a dichotomous variable (case 2b above). If both types of studies are to be combined in the same analysis, one needs to compute the same measure across all studies.
For this, one may need to transform standardized mean differences into log odds ratios (e.g., Cox & Snell, 1989; Chinn, 2000; Hasselblad & Hedges, 1995; Sánchez-Meca et al., 2003). Here, the data need to be specified as described under (1a) and the options for the measure argument are then:
"D2ORN"for the transformed standardized mean difference assuming normal distributions,"D2ORL"for the transformed standardized mean difference assuming logistic distributions.
Both of these transformations provide an estimate of the log odds ratio, the first assuming that the responses within the two groups are normally distributed, while the second assumes that the responses follow logistic distributions.
Alternatively, assuming that the dichotomous outcome in a \(2 \times 2\) table is actually a dichotomized version of the responses on an underlying quantitative scale, it is also possible to estimate the standardized mean difference based on \(2 \times 2\) table data, using either the probit transformed risk difference or a transformation of the odds ratio (e.g., Cox & Snell, 1989; Chinn, 2000; Hasselblad & Hedges, 1995; Sánchez-Meca et al., 2003). Here, the data need to be specified as described under (1b) and the options for the measure argument are then:
"PBIT"for the probit transformed risk difference,"OR2DN"for the transformed odds ratio assuming normal distributions,"OR2DL"for the transformed odds ratio assuming logistic distributions.
All of these transformations provide an estimate of the standardized mean difference, the first two assuming that the responses on the underlying quantitative scale are normally distributed, while the third assumes that the responses follow logistic distributions.
A dataset illustrating the combined analysis of standardized mean differences and probit transformed risk differences is provided in dat.gibson2002.
(2) Measures for Variable Association
Meta-analyses are often used to synthesize studies that examine the direction and strength of the association between two variables measured concurrently and/or without manipulation by experimenters. In this section, a variety of measures will be discussed that may be suitable for a meta-analysis with this purpose. We can distinguish between measures that are applicable when both variables are measured on quantitative scales, when both variables measured are dichotomous, and when the two variables are of mixed types.
(2a) Measures for Two Quantitative Variables
The (Pearson or product-moment) correlation coefficient quantifies the direction and strength of the (linear) relationship between two quantitative variables and is therefore frequently used in meta-analyses. Two alternative measures are a bias-corrected version of the correlation coefficient and Fisher's r-to-z transformed correlation coefficient.
For these measures, one needs to specify ri, the vector with the raw correlation coefficients, and ni, the corresponding sample sizes. The options for the measure argument are then:
"COR"for the raw correlation coefficient,"UCOR"for the raw correlation coefficient corrected for its slight negative bias (based on equation 2.3 in Olkin & Pratt, 1958),"ZCOR"for Fisher's r-to-z transformed correlation coefficient (Fisher, 1921).
If the correlation coefficient is unknown for some studies, but the t-statistics (i.e., \(t_i = r_i \sqrt{n_i - 2} / \sqrt{1 - r_i^2}\)) are available for those studies (for the standard test of \(\text{H}_0{:}\; \rho_i = 0\)), one can specify those values via argument ti, which are then transformed into the corresponding correlation coefficients within the function (the sign of the t-statistics is then taken to be the sign of the correlations). If only the two-sided p-values corresponding to the t-tests are known, one can specify those values via argument pi. However, since a two-sided p-value does not carry information about the sign of the test statistic (and hence neither about the correlation), the sign of the p-values (which can be negative) is used as the sign of the correlation coefficients (e.g., escalc(measure="COR", pi=-0.07, ni=30) yields a negative correlation of -0.3354).
For measure="COR" and measure="UCOR", one can choose between vtype="LS" (the default) for the usual large-sample approximation to compute the sampling variances (i.e., plugging the (biased-corrected) correlation coefficients into equation 12.27 in Borenstein, 2009) and vtype="AV" to compute the sampling variances with the usual large-sample approximation but plugging the sample-size weighted average of the (bias-corrected) correlation coefficients into the equation. For measure="COR", one can also choose vtype="H0" to compute the sampling variances under the null hypothesis (that the true correlations are equal to zero). For measure="UCOR", one can also choose vtype="UB" to compute unbiased estimates of the sampling variances (see Hedges, 1989, but using the exact equation instead of the approximation).
Datasets corresponding to data of this type are provided in dat.mcdaniel1994 and dat.molloy2014.
For meta-analyses involving multiple (dependent) correlation coefficients extracted from the same sample, see also the rcalc function.
(2b) Measures for Two Dichotomous Variables
When the goal of a meta-analysis is to examine the relationship between two dichotomous variables, the data for each study can again be presented in the form of a \(2 \times 2\) table, except that there may not be a clear distinction between the grouping variable and the outcome variable. Moreover, the table may be a result of cross-sectional (i.e., multinomial) sampling, where none of the table margins (except the total sample size) are fixed by the study design.
In particular, assume that the data of interest for a particular study are of the form:
| variable 2, outcome + | variable 2, outcome - | total | ||||
| variable 1, outcome + | ai | bi | n1i | |||
| variable 1, outcome - | ci | di | n2i |
where ai, bi, ci, and di denote the cell frequencies (i.e., the number of individuals falling into a particular category) and n1i and n2i are the row totals.
The phi coefficient and the odds ratio are commonly used measures of association for \(2 \times 2\) table data (e.g., Fleiss & Berlin, 2009). The latter is particularly advantageous, as it is directly comparable to values obtained from stratified sampling (as described earlier). Yule's Q and Yule's Y (Yule, 1912) are additional measures of association for \(2 \times 2\) table data (although they are not typically used in meta-analyses). Finally, assuming that the two dichotomous variables are actually dichotomized versions of the responses on two underlying quantitative scales (and assuming that the two variables follow a bivariate normal distribution), it is also possible to estimate the correlation between the two quantitative variables using the tetrachoric correlation coefficient (Pearson, 1900; Kirk, 1973).
For any of these measures, one needs to specify the cell frequencies via the ai, bi, ci, and di arguments (or alternatively, one can use the ai, ci, n1i, and n2i arguments). The options for the measure argument are then:
"OR"for the log odds ratio,"PHI"for the phi coefficient,"YUQ"for Yule's Q (Yule, 1912),"YUY"for Yule's Y (Yule, 1912),"RTET"for the tetrachoric correlation coefficient.
There are also measures "ZPHI" and "ZTET" for applying Fisher's r-to-z transformation to these measures. This may be useful when combining these with other types of correlation coefficients that were r-to-z transformed. However, note that the r-to-z transformation is not a variance-stabilizing transformation for these measures.
Tables with one or more zero counts are handled as described earlier. For measure="PHI", one must indicate via vtype="ST" or vtype="CS" whether the data for the studies were obtained using stratified or cross-sectional (i.e., multinomial) sampling, respectively (it is also possible to specify an entire vector for the vtype argument in case the sampling scheme differed for the various studies). Note that vtype="LS" is treated like vtype="CS".
A dataset corresponding to data of this type is provided in dat.bourassa1996.
(2d) Measures for Mixed Variable Types
We can also consider measures that can be used to describe the relationship between two variables, where one variable is dichotomous and the other variable measures some quantitative characteristic. In that case, it is likely that study authors again report summary statistics, such as the mean and standard deviation of the measurements within the two groups (defined by the dichotomous variable). Based on this information, one can compute the point-biserial correlation coefficient (Tate, 1954) as a measure of association between the two variables. If the dichotomous variable is actually a dichotomized version of the responses on an underlying quantitative scale (and assuming that the two variables follow a bivariate normal distribution), it is also possible to estimate the correlation between the two variables using the biserial correlation coefficient (Pearson, 1909; Soper, 1914; Jacobs & Viechtbauer, 2017).
Here, one again needs to specify m1i and m2i for the observed means of the two groups, sd1i and sd2i for the observed standard deviations, and n1i and n2i for the number of individuals in each group. The options for the measure argument are then:
"RPB"for the point-biserial correlation coefficient,"RBIS"for the biserial correlation coefficient.
There are also measures "ZPB" and "ZBIS" for applying Fisher's r-to-z transformation to these measures. This may be useful when combining these with other types of correlation coefficients that were r-to-z transformed. However, note that the r-to-z transformation is not a variance-stabilizing transformation for these measures.
If the means and standard deviations are unknown for some studies, one can also use arguments di, ri, ti, or pi to specify standardized mean differences (Cohen's d values), point-biserial correlations, t-statistics from an independent samples t-test, or signed p-values for the t-test, respectively, as described earlier under (1a) (together with the group sizes, these are sufficient statistics for computing the (point-)biserial correlation coefficients).
For measure="RPB", one must indicate via vtype="ST" or vtype="CS" whether the data for the studies were obtained using stratified or cross-sectional (i.e., multinomial) sampling, respectively (it is also possible to specify an entire vector for the vtype argument in case the sampling scheme differed for the various studies). Note that vtype="LS" is treated like vtype="ST".
(3) Measures for Individual Groups
In this section, measures will be described which may be useful when the goal of a meta-analysis is to synthesize studies that characterize some property of individual groups. We will again distinguish between measures that are applicable when the characteristic assessed is a quantitative variable, a dichotomous variable, or when the characteristic represents an event count.
(3a) Measures for Quantitative Variables
The goal of a meta-analysis may be to characterize individual groups, where the response, characteristic, or dependent variable assessed in the individual studies is measured on some quantitative scale. In the simplest case, the raw mean for the quantitative variable is reported for each group, which then becomes the estimate of interest for the meta-analysis. Here, one needs to specify mi, sdi, and ni for the observed means, the observed standard deviations, and the sample sizes, respectively. One can also compute the ‘single-group standardized mean’, where the mean is divided by the standard deviation (when first subtracting some fixed constant from each mean, then this is the ‘single-group standardized mean difference’). If focus is solely on the variability of the measurements, then the log transformed standard deviation is a useful measure (Nakagawa et al., 2015; Raudenbush & Bryk, 1987). For this measure, one only needs to specify sdi and ni. For ratio scale measurements, the log transformed mean or the log transformed coefficient of variation may also be of interest (Nakagawa et al., 2015).
The options for the measure argument are:
"MN"for the raw mean,"SMN"for the single-group standardized mean (difference),"MNLN"for the log transformed mean,"SDLN"for the log transformed standard deviation,"CVLN"for the log transformed coefficient of variation.
Note that sdi is used to specify the standard deviations of the observed values of the response, characteristic, or dependent variable and not the standard errors of the means. Also, the sampling variances for measure="CVLN" are computed as given by equation 27 in Nakagawa et al. (2015), but without the ‘\(-2 \rho \ldots\)’ term, since for normally distributed data (which we assume here) the mean and variance (and transformations thereof) are independent.
(3b) Measures for Dichotomous Variables
A meta-analysis may also be conducted to aggregate studies that provide data about individual groups with respect to a dichotomous dependent variable. Here, one needs to specify xi and ni, denoting the number of individuals experiencing the event of interest and the total number of individuals within each study, respectively. Instead of specifying ni, one can use mi to specify the number of individuals that do not experience the event of interest (i.e., mi=ni-xi). The options for the measure argument are then:
"PR"for the raw proportion,"PLN"for the log transformed proportion,"PLO"for the logit transformed proportion (i.e., log odds),"PRZ"for the probit transformed proportion,"PAS"for the arcsine square root transformed proportion (i.e., the angular transformation),"PFT"for the Freeman-Tukey double arcsine transformed proportion (Freeman & Tukey, 1950).
However, for reasons discussed in Schwarzer et al. (2019) and Röver and Friede (2022), the use of double arcsine transformed proportions for a meta-analysis is not recommended.
Zero cell entries can be problematic for certain measures. When to="only0" (the default), the value of add (the default is 1/2; but see ‘Note’) is added to xi and mi only for studies where xi or mi is equal to 0. When to="all", the value of add is added to xi and mi in all studies. When to="if0all", the value of add is added in all studies, but only when there is at least one study with a zero value for xi or mi. Setting to="none" or add=0 has the same effect: No adjustment to the observed values is made. Depending on the measure and the data, this may lead to division by zero (when this occurs, the resulting value is recoded to NA).
Datasets corresponding to data of this type are provided in dat.pritz1997, dat.debruin2009, dat.hannum2020, and dat.crisafulli2020.
(3c) Measures for Event Counts
Various measures can be used to characterize individual groups when the dependent variable assessed is an event count. Here, one needs to specify xi and ti, denoting the number of events that occurred and the total person-times at risk, respectively. The options for the measure argument are then:
"IR"for the raw incidence rate,"IRLN"for the log transformed incidence rate,"IRS"for the square root transformed incidence rate,"IRFT"for the Freeman-Tukey transformed incidence rate (Freeman & Tukey, 1950).
Measures "IR" and "IRLN" can also be used when meta-analyzing standardized incidence ratios (SIRs), where the observed number of events is divided by the expected number of events. In this case, arguments xi and ti are used to specify the observed and expected number of events in the studies. Since SIRs are not symmetric around 1, it is usually more appropriate to meta-analyze the log transformed SIRs (i.e., using measure "IRLN"), which are symmetric around 0.
Studies with zero events can be problematic, especially for the log transformed incidence rate. Adding a small constant to the number of events is a common solution to this problem. When to="only0" (the default), the value of add (the default is 1/2; but see ‘Note’) is added to xi only in the studies that have zero events. When to="all", the value of add is added to xi in all studies. When to="if0all", the value of add is added to xi in all studies, but only when there is at least one study with zero events. Setting to="none" or add=0 has the same effect: No adjustment to the observed number of events is made. Depending on the measure and the data, this may lead to division by zero (when this occurs, the resulting value is recoded to NA).
(4) Measures for Change or Matched Pairs
The purpose of a meta-analysis may be to assess the amount of change within individual groups (e.g., before and after a treatment or under two different treatments) or when dealing with matched pairs designs.
(4a) Measures for Quantitative Variables
When the response or dependent variable assessed in the individual studies is measured on some quantitative scale, the raw mean change, standardized versions thereof, the common language effect size (area under the curve), or the (log transformed) ratio of means (log response ratio) can be used as measures (Becker, 1988; Gibbons et al., 1993; Lajeunesse, 2011; Morris, 2000). Here, one needs to specify m1i and m2i, the observed means at the two measurement occasions, sd1i and sd2i for the corresponding observed standard deviations, ri for the correlation between the measurements at the two measurement occasions, and ni for the sample size. The options for the measure argument are then:
"MC"for the raw mean change,"SMCC"for the standardized mean change using change score standardization (Gibbons et al., 1993),"SMCR"for the standardized mean change using raw score standardization (Becker, 1988),"SMCRH"for the standardized mean change using raw score standardization with heteroscedastic population variances at the two measurement occasions (Bonett, 2008),"SMCRP"for the standardized mean change using raw score standardization with pooled standard deviations (Cousineau, 2020),"SMCRPH"for the standardized mean change using raw score standardization with pooled standard deviations and heteroscedastic population variances at the two measurement occasions (Bonett, 2008),"CLESCN"(or"AUCCN") for the common language effect size (area under the curve) based on a bivariate normal model for dependent samples,"ROMC"for the log transformed ratio of means (Lajeunesse, 2011).
The raw mean change is simply \(\text{m1i}-\text{m2i}\), while the standardized mean change is given by \((\text{m1i}-\text{m2i})/\text{sdi}\). For measure="SMCC", \(\text{sdi} = \sqrt{\text{sd1i}^2 + \text{sd2i}^2 - 2\times\text{ri}\times\text{sd1i}\times\text{sd2i}}\) is the standard deviation of the change scores, for measure="SMCR" and measure="SMCRH", \(\text{sdi} = \text{sd1i}\), and for measure="SMCRP" and measure="SMCRPH", \(\text{sdi} = \sqrt{\frac{\text{sd1i}^2 + \text{sd2i}^2}{2}}\) is the square root of the average variance. See also Morris and DeShon (2002) for a thorough discussion of the difference between the "SMCC" and "SMCR" change score measures. The log transformed ratio of means is simply \(\log(\text{m1i}/\text{m2i})\)). Bias corrections are applied to measures "SMCC", "SMCR", "SMCRH", "SMCRP", "SMCRPH", and "ROMC" unless correct=FALSE. All of these measures are also applicable for matched pairs designs (subscripts 1 and 2 then simply denote the first and second group that are formed by the matching).
In practice, one often has a mix of information available from the individual studies to compute these measures. In particular, if m1i and m2i are unknown, but the raw mean change is directly reported in a particular study, then one can set m1i to that value and m2i to 0 (making sure that the raw mean change was computed as m1i-m2i within that study and not the other way around). Also, for measures "MC" and "SMCC", if sd1i, sd2i, and ri are unknown, but the standard deviation of the change scores is directly reported, then one can set sd1i to that value and both sd2i and ri to 0. For measure "SMCR", argument sd2i is actually not needed, as the standardization is only based on sd1i (Becker, 1988; Morris, 2000), which is usually the pre-test standard deviation (if the post-test standard deviation should be used, then set sd1i to that). Finally, for measure="SMCC", one can also directly specify standardized mean change values via argument di or the t-statistics from a paired samples t-test or the corresponding two-sided p-values via argument ti or pi, respectively (which are then transformed into the corresponding standardized mean change values within the function). The sign of the p-values (which can be negative) is used as the sign of the standardized mean change values (e.g., escalc(measure="SMCC", pi=-0.018, ni=50) yields a negative standardized mean change value of -0.3408).
Finally, interest may also be focused on differences in the variability of the measurements at the two measurement occasions (or between the two matched groups). For this, the (log transformed) ratio of the standard deviations (also called the ‘variability ratio’) can be used (Senior et al., 2020). To also account for differences in mean levels, the (log transformed) ratio between the coefficients of variation from the two groups (also called the ‘coefficient of variation ratio’) can be a useful measure (Senior et al., 2020). The options for the measure argument are:
"VRC"for the log transformed variability ratio,"CVRC"for the log transformed coefficient of variation ratio.
Note that a bias correction is applied to measure "VRC" unless correct=FALSE. When vtype="LS" (the default), then the sampling variances are computed with equations 21 and 23 from Senior et al. (2020) for "VR" and "CVR", respectively. When vtype="LS2", then equations 22 and 24 based on the second-order Taylor expansions are used instead.
(4b) Measures for Dichotomous Variables
The data for a study examining change in a dichotomous variable gives rise to a paired \(2 \times 2\) table, which is of the form:
| trt 2 outcome 1 | trt 2 outcome 2 | |||
| trt 1 outcome 1 | ai | bi | ||
| trt 1 outcome 2 | ci | di |
where ai, bi, ci, and di denote the cell frequencies. Note that ‘trt1’ and ‘trt2’ may be applied to a single group of subjects or to matched pairs of subjects. Also, ‘trt1’ and ‘trt2’ might refer to two different time points (e.g., before and after a treatment). In any case, the data from such a study can be rearranged into a marginal table of the form:
| outcome 1 | outcome 2 | |||
| trt 1 | ai+bi | ci+di | ||
| trt 2 | ai+ci | bi+di |
which is of the same form as a \(2 \times 2\) table that would arise in a study comparing/contrasting two independent groups.
The options for the measure argument that will compute estimates based on the marginal table are:
"MPRR"for the matched pairs marginal log risk ratio,"MPOR"for the matched pairs marginal log odds ratio,"MPRD"for the matched pairs marginal risk difference.
See Becker and Balagtas (1993), Curtin et al. (2002), Elbourne et al. (2002), Fagerland et al. (2014), May and Johnson (1997), Newcombe (1998), Stedman et al. (2011), and Zou (2007) for discussions of these measures.
The options for the measure argument that will compute estimates based on the paired table are:
"MPORC"for the conditional log odds ratio,"MPPETO"for the conditional log odds ratio estimated with Peto's method.
See Curtin et al. (2002) and Zou (2007) for discussions of these measures.
If only marginal tables are available, then another possibility is to compute the marginal log odds ratios based on these table directly. However, for the correct computation of the sampling variances, the correlations (phi coefficients) from the paired tables must be known (or ‘guestimated’). To use this approach, set measure="MPORM" and use argument ri to specify the correlation coefficients. Instead of specifying ri, one can use argument pi to specify the proportions (or ‘guestimates’ thereof) of individuals (or pairs) that experienced the outcome of interest (i.e., ‘outcome1’ in the paired \(2 \times 2\) table) under both treatments (i.e., pi=ai/(ai+bi+ci+di)). Based on these proportions, the correlation coefficients are then back-calculated and used to compute the correct sampling variances. Note that the values in the marginal tables put constraints on the possible values for ri and pi. If a specified value for ri or pi is not feasible under a given table, the corresponding sampling variance will be NA.
(5) Other Measures for Meta-Analyses
Other measures are sometimes used for meta-analyses that do not directly fall into the categories above. These are described in this section.
Cronbach's alpha and Transformations Thereof
Meta-analytic methods can also be used to aggregate Cronbach's alpha values from multiple studies. This is usually referred to as a ‘reliability generalization meta-analysis’ (Vacha-Haase, 1998). Here, one needs to specify ai, mi, and ni for the observed alpha values, the number of items/replications/parts of the measurement instrument, and the sample sizes, respectively. One can either directly analyze the raw Cronbach's alpha values or transformations thereof (Bonett, 2002, 2010; Hakstian & Whalen, 1976). The options for the measure argument are then:
"ARAW"for raw alpha values,"AHW"for transformed alpha values (Hakstian & Whalen, 1976),"ABT"for transformed alpha values (Bonett, 2002).
Note that the transformations implemented here are slightly different from the ones described by Hakstian and Whalen (1976) and Bonett (2002). In particular, for "AHW", the transformation \(1-(1-\text{ai})^{1/3}\) is used, while for "ABT", the transformation \(-\log(1-\text{ai})\) is used. This ensures that the transformed values are monotonically increasing functions of \(\text{ai}\).
A dataset corresponding to data of this type is provided in dat.bonett2010.
Partial and Semi-Partial Correlations
Aloe and Becker (2012), Aloe and Thompson (2013), and Aloe (2014) describe the use of partial and semi-partial correlation coefficients for meta-analyzing the results from regression models (when the focus is on a common regression coefficient of interest across studies). To compute these measures, one needs to specify ti for the test statistics (i.e., t-tests) of the ‘focal’ regression coefficient of interest, ni for the sample sizes of the studies, mi for the total number of predictors in the regression models (counting the focal predictor of interest, but not the intercept term), and r2i for the \(R^2\) values of the regression models (the latter is only needed when measure="SPCOR" or measure="ZSPCOR"). The options for the measure argument are then:
"PCOR"for the partial correlation coefficient,"ZPCOR"for Fisher's r-to-z transformed partial correlation coefficient,"SPCOR"for the semi-partial correlation coefficient,"ZSPCOR"for Fisher's r-to-z transformed semi-partial correlation coefficient.
Note that the signs of the (semi-)partial correlation coefficients is determined based on the signs of the values specified via the ti argument. Also, while the Fisher transformation can be applied to both measures, it is only a variance-stabilizing transformation for partial correlation coefficients.
If the test statistic (i.e., t-test) of the regression coefficient of interest is unknown for some studies, but the two-sided p-values corresponding to the t-tests are known, one can specify those values via argument pi. However, since a two-sided p-value does not carry information about the sign of the test statistic (and hence neither about the correlation), the sign of the p-values (which can be negative) is used as the sign of the correlation coefficients (e.g., escalc(measure="PCOR", pi=-0.07, mi=5, ni=30) yields a negative partial correlation of -0.3610).
In the rare case that the (semi-)partial correlations are known for some of the studies, then these can be directly specified via the ri argument. This can be useful, for example, when \(\eta^2_p\) (i.e., partial eta squared) is known for the regression coefficient of interest, since the square root thereof is identical to the absolute value of the partial correlation (although the correct sign then still needs to be reconstructed based on other information).
A dataset corresponding to data of this type is provided in dat.aloe2013.
Coefficients of Determination
One can in principle also meta-analyze coefficients of determination (i.e., \(R^2\) values / R-squared values) obtained from a series of linear regression models (however, see the caveat mentioned below). For this, one needs to specify r2i for the \(R^2\) values of the regression models, ni for the sample sizes of the studies, and mi for the number of predictors in the regression models (not counting the intercept term). The options for the measure argument are then:
"R2"for the raw coefficient of determination with predictor values treated as random,"ZR2"for the corresponding r-to-z transformed coefficient of determination,"R2F"for the raw coefficient of determination with predictor values treated as fixed,"ZR2F"for the corresponding r-to-z transformed coefficient of determination.
If the \(R^2\) values are unknown for some studies, but the F-statistics (for the omnibus test of the regression coefficients) are available, one can specify those values via argument fi, which are then transformed into the corresponding \(R^2\) values within the function. If only the p-values corresponding to the F-tests are known, one can specify those values via argument pi (which are then transformed into the F-statistics and then further into the \(R^2\) values).
For measure="R2", one can choose to compute the sampling variances with vtype="LS" (the default) for the large-sample approximation given by equation 27.88 in Kendall and Stuart (1979), vtype="LS2" for the large-sample approximation given by equation 27.87, or vtype="AV" and vtype="AV2" which use the same approximations but plugging the sample-size weighted average of the \(R^2\) values into the equations. For measure="ZR2", the variance-stabilizing transformation \(\frac{1}{2} \log\mathopen{}\left(\frac{1+\sqrt{\text{r2i}}}{1-\sqrt{\text{r2i}}}\right)\mathclose{}\) is used (see Olkin & Finn, 1995, but with the additional \(\frac{1}{2}\) factor), which uses \(1/\text{ni}\) as the large-sample approximation to the sampling variances. The equations used for these measures were derived under the assumption that the values of the outcome variable and the predictors were sampled from a multivariate normal distribution within each study (sometimes called ‘random-X regression’) and that the sample sizes of the studies are large. Moreover, the equations assume that the true \(R^2\) values are non-zero.
For the case where the predictor values are treated as fixed (sometimes called ‘fixed-X regression’), one can use measures "R2F" and "ZR2F". Here, the sampling variances of the \(R^2\) values are computed based on the known relationship between the non-central F-distribution and its non-centrality parameter (which in turn is a function of the true \(R^2\)). However, note that the r-to-z transformation is not a variance-stabilizing transformation for this case.
Given that observed \(R^2\) values cannot be negative, there is no possibility for values to cancel each other out and hence it is guaranteed that the pooled estimate is positive. Hence, a meta-analysis of \(R^2\) values cannot be used to test if the pooled estimate is different from zero (it is by construction as long as the number of studies is sufficiently large).
Relative Excess Heterozygosity
Ziegler et al. (2011) describe the use of meta-analytic methods to examine deviations from the Hardy-Weinberg equilibrium across multiple studies. The relative excess heterozygosity (REH) is the proposed measure for such a meta-analysis, which can be computed by setting measure="REH". Here, one needs to specify ai for the number of individuals with homozygous dominant alleles, bi for the number of individuals with heterozygous alleles, and ci for the number of individuals with homozygous recessives alleles.
Note that the log is taken of the REH values, which makes this measure symmetric around 0 and results in a sampling distribution that is closer to normality.
A dataset corresponding to data of this type is provided in dat.frank2008.
(6) Converting a Data Frame to an 'escalc' Object
The function can also be used to convert a regular data frame to an ‘escalc’ object. One simply sets the measure argument to one of the options described above (or to measure="GEN" for a set of generic estimates not further specified) and passes the estimates via the yi argument and the corresponding sampling variances via the vi argument (or the standard errors via the sei argument) to the function.
Flagging Large Estimates
For some of the measures above, the function runs a check to examine if one or more of the computed estimates are unusually large. This may happen due to data extraction or reporting errors and might warrant double-checking the values used as input to the function.
At the moment, this check is conducted for standardized mean differences (i.e., measure "SMD") and its variants. A common mistake when computing such measures is that standard errors of the means are used in place of standard deviations. Recall that a standardized mean difference (before the bias correction) is given by \(d = (\text{m1i}-\text{m2i})/\text{sdi}\), where \(\text{sdi} = \sqrt{\frac{(\text{n1i}-1)\text{sd1i}^2 + (\text{n2i}-1)\text{sd2i}^2}{\text{n1i}+\text{n2i}-2}}\) is the pooled standard deviation and sd1i and sd2i are the standard deviations of the two groups (see section (1a) above). When using the standard errors of the means in place of the standard deviations, then sdi will be too small (since the standard error of a mean is equal to the standard deviation divided by the square root of the sample size) and the computed \(d\) value will be too large. An excessively large \(d\) value might be an indication that such a mistake was made. Since this invalidates all meta-analytic results computed based on such a value, the function checks if any of the calculated standardized mean differences are larger than 2 in absolute value. This cutoff is somewhat arbitrary and can be adjusted via the cutoff argument. The check can be skipped by setting cutoff=NA. Also, to avoid annoyance due to false positives, the check is only conducted once for a given set of estimates within a session.
Other Arguments
Argument slab can be used to specify (study) labels for the estimates. These labels are passed on to other functions and used as needed (e.g., for labeling the estimates in a forest plot). Note that missing values in the labels are not allowed.
The flip argument, when set to TRUE, can be used to flip the sign of the estimates. This can also be a logical vector to indicate for which studies the sign should be flipped. The argument can also be a numeric vector to specify a multiplier for the estimates (the corresponding sampling variances are adjusted accordingly).
The subset argument can be used to select the studies that will be included in the data frame returned by the function. On the other hand, the include argument simply selects for which studies the measure will be computed (if it shouldn't be computed for all of them).
Value
An object of class c("escalc","data.frame"). The object is a data frame containing the following components:
- yi
vector with the estimates.
- vi
vector with the corresponding sampling variances.
If a data frame was specified via the data argument and append=TRUE, then variables yi and vi are appended to this data frame. Note that the var.names argument actually specifies the names of these two variables ("yi" and "vi" are the defaults).
If the data frame already contains two variables with names as specified by the var.names argument, the values for these two variables will be overwritten when replace=TRUE (which is the default). By setting replace=FALSE, only values that are NA will be replaced.
The object is formatted and printed with the print function. The summary function can be used to obtain confidence intervals around the estimates. See methods.escalc for some additional method functions for "escalc" objects.
With the aggregate function, one can aggregate multiple estimates belonging to the same study (or some other clustering variable) into a single combined estimate.
Note
The variable names specified under var.names should be syntactically valid variable names. If necessary, they are adjusted so that they are.
Although the default value for add is 1/2, for certain measures the use of such a bias correction makes little sense and for these measures, the function internally sets add=0. This applies to the following measures: "AS", "PHI", "ZPHI", "RTET", "ZTET", "IRSD", "PAS", "PFT", "IRS", and "IRFT". One can still force the use of the bias correction by explicitly setting the add argument to some non-zero value when calling the function.
References
Aloe, A. M. (2014). An empirical investigation of partial effect sizes in meta-analysis of correlational data. Journal of General Psychology, 141(1), 47–64. https://doi.org/10.1080/00221309.2013.853021
Aloe, A. M., & Becker, B. J. (2012). An effect size for regression predictors in meta-analysis. Journal of Educational and Behavioral Statistics, 37(2), 278–297. https://doi.org/10.3102/1076998610396901
Aloe, A. M., & Thompson, C. G. (2013). The synthesis of partial effect sizes. Journal of the Society for Social Work and Research, 4(4), 390–405. https://doi.org/10.5243/jsswr.2013.24
Bagos, P. G., & Nikolopoulos, G. K. (2009). Mixed-effects Poisson regression models for meta-analysis of follow-up studies with constant or varying durations. The International Journal of Biostatistics, 5(1). https://doi.org/10.2202/1557-4679.1168
Becker, B. J. (1988). Synthesizing standardized mean-change measures. British Journal of Mathematical and Statistical Psychology, 41(2), 257–278. https://doi.org/10.1111/j.2044-8317.1988.tb00901.x
Becker, M. P., & Balagtas, C. C. (1993). Marginal modeling of binary cross-over data. Biometrics, 49(4), 997–1009. https://doi.org/10.2307/2532242
Bonett, D. G. (2002). Sample size requirements for testing and estimating coefficient alpha. Journal of Educational and Behavioral Statistics, 27(4), 335–340. https://doi.org/10.3102/10769986027004335
Bonett, D. G. (2008). Confidence intervals for standardized linear contrasts of means. Psychological Methods, 13(2), 99–109. https://doi.org/10.1037/1082-989X.13.2.99
Bonett, D. G. (2009). Meta-analytic interval estimation for standardized and unstandardized mean differences. Psychological Methods, 14(3), 225–238. https://doi.org/10.1037/a0016619
Bonett, D. G. (2010). Varying coefficient meta-analytic methods for alpha reliability. Psychological Methods, 15(4), 368–385. https://doi.org/10.1037/a0020142
Borenstein, M. (2009). Effect sizes for continuous data. In H. Cooper, L. V. Hedges, & J. C. Valentine (Eds.), The handbook of research synthesis and meta-analysis (2nd ed., pp. 221–235). New York: Russell Sage Foundation.
Chinn, S. (2000). A simple method for converting an odds ratio to effect size for use in meta-analysis. Statistics in Medicine, 19(22), 3127–3131. https://doi.org/10.1002/1097-0258(20001130)19:22<3127::aid-sim784>3.0.co;2-m
Cho, H., Matthews, G. J., & Harel, O. (2019). Confidence intervals for the area under the receiver operating characteristic curve in the presence of ignorable missing data. International Statistical Review, 87(1), 152–177. https://doi.org/10.1111/insr.12277
Cohen, J. (1988). Statistical power analysis for the behavioral sciences (2nd ed.). Hillsdale, NJ: Lawrence Erlbaum Associates.
Cousineau, D. (2020). Approximating the distribution of Cohen's d_p in within-subject designs. The Quantitative Methods for Psychology, 16(4), 418–421. https://doi.org/10.20982/tqmp.16.4.p418
Cox, D. R., & Snell, E. J. (1989). Analysis of binary data (2nd ed.). London: Chapman & Hall.
Curtin, F., Elbourne, D., & Altman, D. G. (2002). Meta-analysis combining parallel and cross-over clinical trials. II: Binary outcomes. Statistics in Medicine, 21(15), 2145–2159. https://doi.org/10.1002/sim.1206
Elbourne, D. R., Altman, D. G., Higgins, J. P. T., Curtin, F., Worthington, H. V., & Vail, A. (2002). Meta-analyses involving cross-over trials: Methodological issues. International Journal of Epidemiology, 31(1), 140–149. https://doi.org/10.1093/ije/31.1.140
Fagerland, M. W., Lydersen, S., & Laake, P. (2014). Recommended tests and confidence intervals for paired binomial proportions. Statistics in Medicine, 33(16), 2850–2875. https://doi.org/10.1002/sim.6148
Fisher, R. A. (1921). On the “probable error” of a coefficient of correlation deduced from a small sample. Metron, 1, 1–32. https://hdl.handle.net/2440/15169
Fleiss, J. L., & Berlin, J. (2009). Effect sizes for dichotomous data. In H. Cooper, L. V. Hedges, & J. C. Valentine (Eds.), The handbook of research synthesis and meta-analysis (2nd ed., pp. 237–253). New York: Russell Sage Foundation.
Freeman, M. F., & Tukey, J. W. (1950). Transformations related to the angular and the square root. Annals of Mathematical Statistics, 21(4), 607–611. https://doi.org/10.1214/aoms/1177729756
Gibbons, R. D., Hedeker, D. R., & Davis, J. M. (1993). Estimation of effect size from a series of experiments involving paired comparisons. Journal of Educational Statistics, 18(3), 271–279. https://doi.org/10.3102/10769986018003271
Goddard, M. J., & Hinberg, I. (1990). Receiver operator characteristic (ROC) curves and non‐normal data: An empirical study. Statistics in Medicine, 9(3), 325–337. https://doi.org/10.1002/sim.4780090315
Hakstian, A. R., & Whalen, T. E. (1976). A k-sample significance test for independent alpha coefficients. Psychometrika, 41(2), 219–231. https://doi.org/10.1007/BF02291840
Hanley, J. A., & McNeil, B. J. (1982). The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology, 143(1), 29–36. https://doi.org/10.1148/radiology.143.1.7063747
Hasselblad, V., & Hedges, L. V. (1995). Meta-analysis of screening and diagnostic tests. Psychological Bulletin, 117(1), 167–178. https://doi.org/10.1037/0033-2909.117.1.167
Hedges, L. V. (1981). Distribution theory for Glass's estimator of effect size and related estimators. Journal of Educational Statistics, 6(2), 107–128. https://doi.org/10.3102/10769986006002107
Hedges, L. V. (1982). Estimation of effect size from a series of independent experiments. Psychological Bulletin, 92(2), 490–499. https://doi.org/10.1037/0033-2909.92.2.490
Hedges, L. V. (1983). A random effects model for effect sizes. Psychological Bulletin, 93(2), 388–395. https://doi.org/10.1037/0033-2909.93.2.388
Hedges, L. V. (1989). An unbiased correction for sampling error in validity generalization studies. Journal of Applied Psychology, 74(3), 469–477. https://doi.org/10.1037/0021-9010.74.3.469
Hedges, L. V., Gurevitch, J., & Curtis, P. S. (1999). The meta-analysis of response ratios in experimental ecology. Ecology, 80(4), 1150–1156. https://doi.org/10.1890/0012-9658(1999)080[1150:TMAORR]2.0.CO;2
Higgins, J. P. T., Thomas, J., Chandler, J., Cumpston, M., Li, T., Page, M. J., & Welch, V. A. (Eds.) (2019). Cochrane handbook for systematic reviews of interventions (2nd ed.). Chichester, UK: Wiley. https://training.cochrane.org/handbook
Jacobs, P., & Viechtbauer, W. (2017). Estimation of the biserial correlation and its sampling variance for use in meta-analysis. Research Synthesis Methods, 8(2), 161–180. https://doi.org/10.1002/jrsm.1218
Kendall, M., & Stuart, A. (1979). Kendall's advanced theory of statistics, Vol. 2: Inference and relationship (4th ed.). New York: Macmillan.
Kirk, D. B. (1973). On the numerical approximation of the bivariate normal (tetrachoric) correlation coefficient. Psychometrika, 38(2), 259–268. https://doi.org/10.1007/BF02291118
Lajeunesse, M. J. (2011). On the meta-analysis of response ratios for studies with correlated and multi-group designs. Ecology, 92(11), 2049–2055. https://doi.org/10.1890/11-0423.1
Lajeunesse, M. J. (2015). Bias and correction for the log response ratio in ecological meta-analysis. Ecology, 96(8), 2056–2063. https://doi.org/10.1890/14-2402.1
May, W. L., & Johnson, W. D. (1997). Confidence intervals for differences in correlated binary proportions. Statistics in Medicine, 16(18), 2127–2136. https://doi.org/10.1002/(SICI)1097-0258(19970930)16:18<2127::AID-SIM633>3.0.CO;2-W
McGraw, K. O., & Wong, S. P. (1992). A common language effect size statistic. Psychological Bulletin, 111(2), 361–365. https://doi.org/10.1037/0033-2909.111.2.361
Morris, S. B. (2000). Distribution of the standardized mean change effect size for meta-analysis on repeated measures. British Journal of Mathematical and Statistical Psychology, 53(1), 17–29. https://doi.org/10.1348/000711000159150
Morris, S. B., & DeShon, R. P. (2002). Combining effect size estimates in meta-analysis with repeated measures and independent-groups designs. Psychological Methods, 7(1), 105–125. https://doi.org/10.1037/1082-989x.7.1.105
Nakagawa, S., Poulin, R., Mengersen, K., Reinhold, K., Engqvist, L., Lagisz, M., & Senior, A. M. (2015). Meta-analysis of variation: Ecological and evolutionary applications and beyond. Methods in Ecology and Evolution, 6(2), 143–152. https://doi.org/10.1111/2041-210x.12309
Nakagawa, S., Noble, D. W. A., Lagisz, M., Spake, R., Viechtbauer, W., & Senior, A. M. (2023). A robust and readily implementable method for the meta-analysis of response ratios with and without missing standard deviations. Ecology Letters, 26(2), 232–244. https://doi.org/10.1111/ele.14144
Newcombe, R. G. (1998). Improved confidence intervals for the difference between binomial proportions based on paired data. Statistics in Medicine, 17(22), 2635–2650. https://doi.org/10.1002/(SICI)1097-0258(19981130)17:22<2635::AID-SIM954>3.0.CO;2-C
Newcombe, R. G. (2006). Confidence intervals for an effect size measure based on the Mann-Whitney statistic. Part 2: Asymptotic methods and evaluation. Statistics in Medicine, 25(4), 559–573. https://doi.org/10.1002/sim.2324
Olkin, I., & Finn, J. D. (1995). Correlations redux. Psychological Bulletin, 118(1), 155–164. https://doi.org/10.1037/0033-2909.118.1.155
Olkin, I., & Pratt, J. W. (1958). Unbiased estimation of certain correlation coefficients. Annals of Mathematical Statistics, 29(1), 201–211. https://doi.org/10.1214/aoms/1177706717
Pearson, K. (1900). Mathematical contributions to the theory of evolution. VII. On the correlation of characters not quantitatively measurable. Philosophical Transactions of the Royal Society of London, Series A, 195, 1–47. https://doi.org/10.1098/rsta.1900.0022
Pearson, K. (1909). On a new method of determining correlation between a measured character A, and a character B, of which only the percentage of cases wherein B exceeds (or falls short of) a given intensity is recorded for each grade of A. Biometrika, 7(1/2), 96–105. https://doi.org/10.1093/biomet/7.1-2.96
Raudenbush, S. W., & Bryk, A. S. (1987). Examining correlates of diversity. Journal of Educational Statistics, 12(3), 241–269. https://doi.org/10.3102/10769986012003241
Rothman, K. J., Greenland, S., & Lash, T. L. (2008). Modern epidemiology (3rd ed.). Philadelphia: Lippincott Williams & Wilkins.
Röver, C., & Friede, T. (2022). Double arcsine transform not appropriate for meta‐analysis. Research Synthesis Methods, 13(5), 645–648. https://doi.org/10.1002/jrsm.1591
Rücker, G., Schwarzer, G., Carpenter, J., & Olkin, I. (2009). Why add anything to nothing? The arcsine difference as a measure of treatment effect in meta-analysis with zero cells. Statistics in Medicine, 28(5), 721–738. https://doi.org/10.1002/sim.3511
Sánchez-Meca, J., Marín-Martínez, F., & Chacón-Moscoso, S. (2003). Effect-size indices for dichotomized outcomes in meta-analysis. Psychological Methods, 8(4), 448–467. https://doi.org/10.1037/1082-989X.8.4.448
Schwarzer, G., Chemaitelly, H., Abu-Raddad, L. J., & Rücker, G. (2019). Seriously misleading results using inverse of Freeman-Tukey double arcsine transformation in meta-analysis of single proportions. Research Synthesis Methods, 10(3), 476–483. https://doi.org/10.1002/jrsm.1348
Senior, A. M., Viechtbauer, W., & Nakagawa, S. (2020). Revisiting and expanding the meta-analysis of variation: The log coefficient of variation ratio. Research Synthesis Methods, 11(4), 553–567. https://doi.org/10.1002/jrsm.1423
Soper, H. E. (1914). On the probable error of the bi-serial expression for the correlation coefficient. Biometrika, 10(2/3), 384–390. https://doi.org/10.1093/biomet/10.2-3.384
Stedman, M. R., Curtin, F., Elbourne, D. R., Kesselheim, A. S., & Brookhart, M. A. (2011). Meta-analyses involving cross-over trials: Methodological issues. International Journal of Epidemiology, 40(6), 1732–1734. https://doi.org/10.1093/ije/dyp345
Tate, R. F. (1954). Correlation between a discrete and a continuous variable: Point-biserial correlation. Annals of Mathematical Statistics, 25(3), 603–607. https://doi.org/10.1214/aoms/1177728730
Vacha-Haase, T. (1998). Reliability generalization: Exploring variance in measurement error affecting score reliability across studies. Educational and Psychological Measurement, 58(1), 6–20. https://doi.org/10.1177/0013164498058001002
Viechtbauer, W. (2010). Conducting meta-analyses in R with the metafor package. Journal of Statistical Software, 36(3), 1–48. https://doi.org/10.18637/jss.v036.i03
Yule, G. U. (1912). On the methods of measuring association between two attributes. Journal of the Royal Statistical Society, 75(6), 579–652. https://doi.org/10.2307/2340126
Yusuf, S., Peto, R., Lewis, J., Collins, R., & Sleight, P. (1985). Beta blockade during and after myocardial infarction: An overview of the randomized trials. Progress in Cardiovascular Disease, 27(5), 335–371. https://doi.org/10.1016/s0033-0620(85)80003-7
Ziegler, A., Steen, K. V. & Wellek, S. (2011). Investigating Hardy-Weinberg equilibrium in case-control or cohort studies or meta-analysis. Breast Cancer Research and Treatment, 128(1), 197–201. https://doi.org/10.1007/s10549-010-1295-z
Zou, G. Y. (2007). One relative risk versus two odds ratios: Implications for meta-analyses involving paired and unpaired binary data. Clinical Trials, 4(1), 25–31. https://doi.org/10.1177/1740774506075667
See also
print and summary for the print and summary methods.
conv.2x2 for a function to reconstruct the cell frequencies of \(2 \times 2\) tables based on other summary statistics.
conv.fivenum for a function to convert five-number summary values to means and standard deviations (needed to compute various effect size measures, such as raw or standardized mean differences and ratios of means / response ratios).
conv.wald for a function to convert Wald-type confidence intervals and test statistics to sampling variances.
conv.delta for a function to transform estimates and their sampling variances using the delta method.
vcalc for a function to construct or approximate the variance-covariance matrix of dependent estimates.
rcalc for a function to construct the variance-covariance matrix of dependent correlation coefficients.
rma.uni and rma.mv for model fitting functions that can take the calculated estimates (and the corresponding sampling variances) as input.
rma.mh, rma.peto, and rma.glmm for model fitting functions that take similar inputs.
Examples
############################################################################
### data from the meta-analysis by Coliditz et al. (1994) on the efficacy of
### BCG vaccine in the prevention of tuberculosis dat.bcg
dat.bcg
#> trial author year tpos tneg cpos cneg ablat alloc
#> 1 1 Aronson 1948 4 119 11 128 44 random
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic
#> 13 13 Comstock et al 1976 27 16886 29 17825 33 systematic
### calculate log risk ratios and corresponding sampling variances
dat <- escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg)
dat
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random -1.6209 0.2230
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random 0.0120 0.0040
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random -0.4694 0.0564
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic -1.3713 0.0730
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic -0.3394 0.0124
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic 0.4459 0.5325
#> 13 13 Comstock et al 1976 27 16886 29 17825 33 systematic -0.0173 0.0714
#>
### suppose that for a particular study, yi and vi are known (i.e., have
### already been calculated) but the 2x2 table counts are not known; with
### replace=FALSE, the yi and vi values for that study are not replaced
dat[1:12,10:11] <- NA
dat[13,4:7] <- NA
dat
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random NA NA
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random NA NA
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random NA NA
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random NA NA
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate NA NA
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate NA NA
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random NA NA
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random NA NA
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random NA NA
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic NA NA
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic NA NA
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic NA NA
#> 13 13 Comstock et al 1976 NA NA NA NA 33 systematic -0.0173 0.0714
#>
dat <- escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat, replace=FALSE)
dat
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random -1.6209 0.2230
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random 0.0120 0.0040
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random -0.4694 0.0564
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic -1.3713 0.0730
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic -0.3394 0.0124
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic 0.4459 0.5325
#> 13 13 Comstock et al 1976 NA NA NA NA 33 systematic -0.0173 0.0714
#>
### illustrate difference between 'subset' and 'include' arguments
escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg, subset=1:6)
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#>
escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg, include=1:6)
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random NA NA
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random NA NA
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random NA NA
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic NA NA
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic NA NA
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic NA NA
#> 13 13 Comstock et al 1976 27 16886 29 17825 33 systematic NA NA
#>
### illustrate the 'var.names' argument
escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg, var.names=c("lnrr","var"))
#>
#> trial author year tpos tneg cpos cneg ablat alloc lnrr var
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random -1.6209 0.2230
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random 0.0120 0.0040
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random -0.4694 0.0564
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic -1.3713 0.0730
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic -0.3394 0.0124
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic 0.4459 0.5325
#> 13 13 Comstock et al 1976 27 16886 29 17825 33 systematic -0.0173 0.0714
#>
### illustrate the 'slab' argument
dat <- escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg,
data=dat.bcg, slab=paste0(author, ", ", year))
dat
#>
#> trial author year tpos tneg cpos cneg ablat alloc yi vi
#> 1 1 Aronson 1948 4 119 11 128 44 random -0.8893 0.3256
#> 2 2 Ferguson & Simes 1949 6 300 29 274 55 random -1.5854 0.1946
#> 3 3 Rosenthal et al 1960 3 228 11 209 42 random -1.3481 0.4154
#> 4 4 Hart & Sutherland 1977 62 13536 248 12619 52 random -1.4416 0.0200
#> 5 5 Frimodt-Moller et al 1973 33 5036 47 5761 13 alternate -0.2175 0.0512
#> 6 6 Stein & Aronson 1953 180 1361 372 1079 44 alternate -0.7861 0.0069
#> 7 7 Vandiviere et al 1973 8 2537 10 619 19 random -1.6209 0.2230
#> 8 8 TPT Madras 1980 505 87886 499 87892 13 random 0.0120 0.0040
#> 9 9 Coetzee & Berjak 1968 29 7470 45 7232 27 random -0.4694 0.0564
#> 10 10 Rosenthal et al 1961 17 1699 65 1600 42 systematic -1.3713 0.0730
#> 11 11 Comstock et al 1974 186 50448 141 27197 18 systematic -0.3394 0.0124
#> 12 12 Comstock & Webster 1969 5 2493 3 2338 33 systematic 0.4459 0.5325
#> 13 13 Comstock et al 1976 27 16886 29 17825 33 systematic -0.0173 0.0714
#>
### note: the output looks the same but the study labels are stored as an attribute with the
### effect size estimates (together with the total sample size of the studies and the chosen
### effect size measure)
dat$yi
#> [1] -0.88931133 -1.58538866 -1.34807315 -1.44155119 -0.21754732 -0.78611559 -1.62089822 0.01195233
#> [9] -0.46941765 -1.37134480 -0.33935883 0.44591340 -0.01731395
#> attr(,"ni")
#> [1] 262 609 451 26465 10877 2992 3174 176782 14776 3381 77972 4839 34767
#> attr(,"slab")
#> [1] "Aronson, 1948" "Ferguson & Simes, 1949" "Rosenthal et al, 1960"
#> [4] "Hart & Sutherland, 1977" "Frimodt-Moller et al, 1973" "Stein & Aronson, 1953"
#> [7] "Vandiviere et al, 1973" "TPT Madras, 1980" "Coetzee & Berjak, 1968"
#> [10] "Rosenthal et al, 1961" "Comstock et al, 1974" "Comstock & Webster, 1969"
#> [13] "Comstock et al, 1976"
#> attr(,"measure")
#> [1] "RR"
### this information can then be used by other functions; for example in a forest plot
forest(dat$yi, dat$vi)
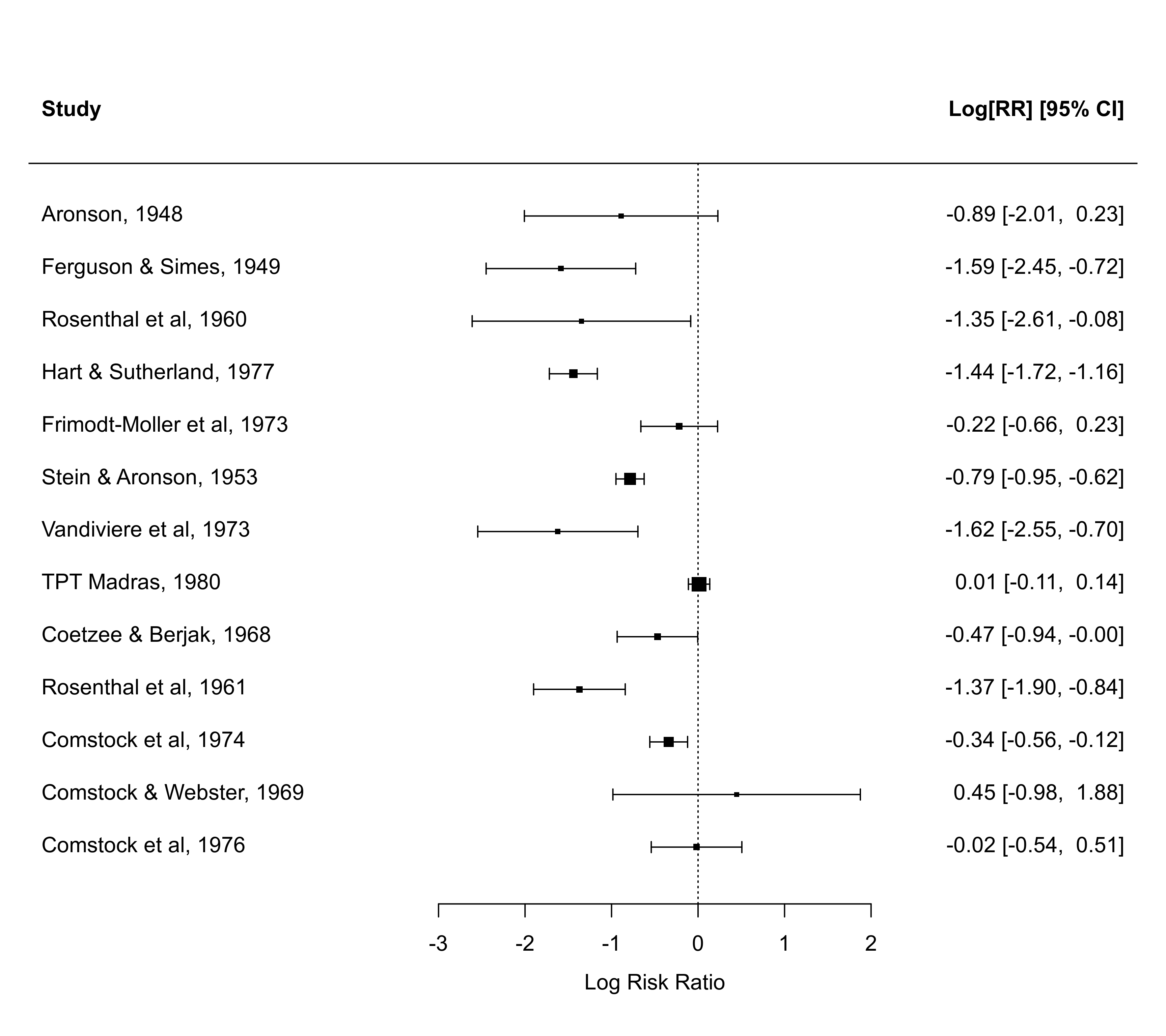 ############################################################################
### convert a regular data frame to an 'escalc' object
### dataset from Lipsey & Wilson (2001), Table 7.1, page 130
dat <- data.frame(id = c(100, 308, 1596, 2479, 9021, 9028, 161, 172, 537, 7049),
yi = c(-0.33, 0.32, 0.39, 0.31, 0.17, 0.64, -0.33, 0.15, -0.02, 0.00),
vi = c(0.084, 0.035, 0.017, 0.034, 0.072, 0.117, 0.102, 0.093, 0.012, 0.067),
random = c(0, 0, 0, 0, 0, 0, 1, 1, 1, 1),
intensity = c(7, 3, 7, 5, 7, 7, 4, 4, 5, 6))
dat
#> id yi vi random intensity
#> 1 100 -0.33 0.084 0 7
#> 2 308 0.32 0.035 0 3
#> 3 1596 0.39 0.017 0 7
#> 4 2479 0.31 0.034 0 5
#> 5 9021 0.17 0.072 0 7
#> 6 9028 0.64 0.117 0 7
#> 7 161 -0.33 0.102 1 4
#> 8 172 0.15 0.093 1 4
#> 9 537 -0.02 0.012 1 5
#> 10 7049 0.00 0.067 1 6
dat <- escalc(measure="SMD", yi=yi, vi=vi, data=dat, slab=paste("Study ID:", id), digits=3)
dat
#>
#> id yi vi random intensity
#> 1 100 -0.330 0.084 0 7
#> 2 308 0.320 0.035 0 3
#> 3 1596 0.390 0.017 0 7
#> 4 2479 0.310 0.034 0 5
#> 5 9021 0.170 0.072 0 7
#> 6 9028 0.640 0.117 0 7
#> 7 161 -0.330 0.102 1 4
#> 8 172 0.150 0.093 1 4
#> 9 537 -0.020 0.012 1 5
#> 10 7049 0.000 0.067 1 6
#>
############################################################################
############################################################################
### convert a regular data frame to an 'escalc' object
### dataset from Lipsey & Wilson (2001), Table 7.1, page 130
dat <- data.frame(id = c(100, 308, 1596, 2479, 9021, 9028, 161, 172, 537, 7049),
yi = c(-0.33, 0.32, 0.39, 0.31, 0.17, 0.64, -0.33, 0.15, -0.02, 0.00),
vi = c(0.084, 0.035, 0.017, 0.034, 0.072, 0.117, 0.102, 0.093, 0.012, 0.067),
random = c(0, 0, 0, 0, 0, 0, 1, 1, 1, 1),
intensity = c(7, 3, 7, 5, 7, 7, 4, 4, 5, 6))
dat
#> id yi vi random intensity
#> 1 100 -0.33 0.084 0 7
#> 2 308 0.32 0.035 0 3
#> 3 1596 0.39 0.017 0 7
#> 4 2479 0.31 0.034 0 5
#> 5 9021 0.17 0.072 0 7
#> 6 9028 0.64 0.117 0 7
#> 7 161 -0.33 0.102 1 4
#> 8 172 0.15 0.093 1 4
#> 9 537 -0.02 0.012 1 5
#> 10 7049 0.00 0.067 1 6
dat <- escalc(measure="SMD", yi=yi, vi=vi, data=dat, slab=paste("Study ID:", id), digits=3)
dat
#>
#> id yi vi random intensity
#> 1 100 -0.330 0.084 0 7
#> 2 308 0.320 0.035 0 3
#> 3 1596 0.390 0.017 0 7
#> 4 2479 0.310 0.034 0 5
#> 5 9021 0.170 0.072 0 7
#> 6 9028 0.640 0.117 0 7
#> 7 161 -0.330 0.102 1 4
#> 8 172 0.150 0.093 1 4
#> 9 537 -0.020 0.012 1 5
#> 10 7049 0.000 0.067 1 6
#>
############################################################################