Forest Plots (Method for 'cumul.rma' Objects)
forest.cumul.rma.RdFunction to create forest plots for objects of class "cumul.rma".
# S3 method for class 'cumul.rma'
forest(x, annotate=TRUE, header=TRUE,
xlim, alim, olim, ylim, at, steps=5,
refline=0, digits=2L, width,
xlab, ilab, ilab.lab, ilab.xpos, ilab.pos,
transf, atransf, targs, rows,
efac=1, pch, psize, col, shade, colshade,
lty, fonts, cex, cex.lab, cex.axis, ...)Arguments
- x
an object of class
"cumul.rma"obtained withcumul.- annotate
logical to specify whether annotations should be added to the plot (the default is
TRUE).- header
logical to specify whether column headings should be added to the plot (the default is
TRUE). Can also be a character vector to specify the left and right headings (or only the left one).- xlim
horizontal limits of the plot region. If unspecified, the function sets the horizontal plot limits to some sensible values.
- alim
the x-axis limits. If unspecified, the function sets the x-axis limits to some sensible values.
- olim
argument to specify observation limits. If unspecified, no limits are used.
- ylim
the y-axis limits of the plot. If unspecified, the function sets the y-axis limits to some sensible values. Can also be a single value to set the lower bound (while the upper bound is still set automatically).
- at
position of the x-axis tick marks and corresponding labels. If unspecified, the function sets the tick mark positions/labels to some sensible values.
- steps
the number of tick marks for the x-axis (the default is 5). Ignored when the positions are specified via the
atargument.- refline
numeric value to specify the location of the vertical ‘reference’ line (the default is 0). The line can be suppressed by setting this argument to
NA. Can also be a vector to add multiple lines.- digits
integer to specify the number of decimal places to which the annotations and tick mark labels of the x-axis should be rounded (the default is
2L). Can also be a vector of two integers, the first to specify the number of decimal places for the annotations, the second for the x-axis labels. When specifying an integer (e.g.,2L), trailing zeros after the decimal mark are dropped for the x-axis labels. When specifying a numeric value (e.g.,2), trailing zeros are retained.- width
optional integer to manually adjust the width of the columns for the annotations (either a single integer or a vector of the same length as the number of annotation columns).
- xlab
title for the x-axis. If unspecified, the function sets an appropriate axis title. Can also be a vector of three/two values (to also/only add labels at the end points of the x-axis limits).
- ilab
optional vector, matrix, or data frame providing additional information about the studies that should be added to the plot.
- ilab.lab
optional character vector with (column) labels for the variable(s) given via
ilab.- ilab.xpos
optional numeric vector to specify the horizontal position(s) of the variable(s) given via
ilab.- ilab.pos
integer(s) (either 1, 2, 3, or 4) to specify the alignment of the variable(s) given via
ilab(2 means right, 4 means left aligned). If unspecified, the default is to center the values.- transf
optional argument to specify a function to transform the estimates and confidence interval bounds (e.g.,
transf=exp; see also transf). If unspecified, no transformation is used.- atransf
optional argument to specify a function to transform the x-axis labels and annotations (e.g.,
atransf=exp; see also transf). If unspecified, no transformation is used.- targs
optional arguments needed by the function specified via
transforatransf.- rows
optional vector to specify the rows (or more generally, the positions) for plotting the estimates. Can also be a single value to specify the row of the first estimate (the remaining estimates are then plotted below this starting row).
- efac
vertical expansion factor for confidence interval limits and arrows. The default value of 1 should usually work fine. Can also be a vector of two numbers, the first for CI limits, the second for arrows.
- pch
plotting symbol to use for the estimates. By default, a filled square is used. See
pointsfor other options. Can also be a vector of values.- psize
numeric value to specify the point sizes for the estimates (the default is 1). Can also be a vector of values.
- col
optional character string to specify the color of the estimates. Can also be a vector.
- shade
optional character string or a (logical or numeric) vector for shading rows of the plot.
- colshade
optional argument to specify the color for the shading.
- lty
optional argument to specify the line type for the confidence intervals. If unspecified, the function sets this to
"solid"by default.- fonts
optional character string to specify the font for the study labels, annotations, and the extra information (if specified via
ilab). If unspecified, the default font is used.- cex
optional character and symbol expansion factor. If unspecified, the function sets this to a sensible value.
- cex.lab
optional expansion factor for the x-axis title. If unspecified, the function sets this to a sensible value.
- cex.axis
optional expansion factor for the x-axis labels. If unspecified, the function sets this to a sensible value.
- ...
other arguments.
Details
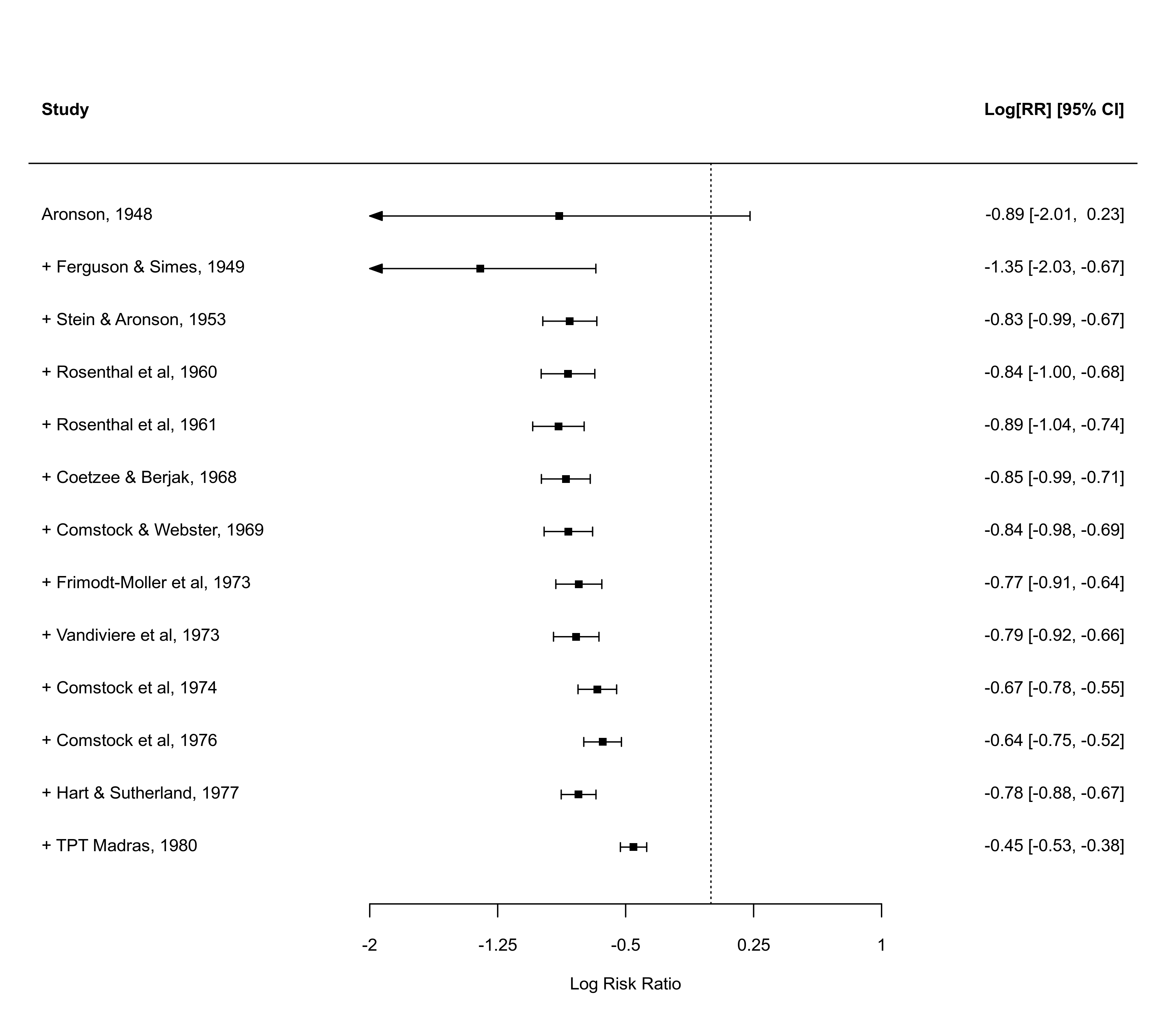
The cumul function can be used to conduct a cumulative meta-analysis. Passing an object created by this function to forest() creates a cumulative forest plot that shows the pooled estimate with corresponding confidence interval bounds as one estimate at a time is added to the analysis.
See forest.default and forest.rma for further details on the purpose of the various arguments.
Note
The function sets some sensible values for the optional arguments, but it may be necessary to adjust these in certain circumstances.
The function actually returns some information about the chosen values invisibly. Printing this information is useful as a starting point to customize the plot.
If the number of studies is quite large, the labels, annotations, and symbols may become quite small and impossible to read. Stretching the plot window vertically may then provide a more readable figure (one should call the function again after adjusting the window size, so that the label/symbol sizes can be properly adjusted). Also, the cex, cex.lab, and cex.axis arguments are then useful to adjust the symbol and text sizes.
If the estimates are bounded (e.g., correlations are bounded between -1 and +1, proportions are bounded between 0 and 1), one can use the olim argument to enforce those limits (i.e., the pooled estimates and confidence interval bounds cannot exceed those limits then).
The lty argument can also be a vector of two elements, the first for specifying the line type of the individual CIs ("solid" by default), the second for the line type of the horizontal line that is automatically added to the plot ("solid" by default; set to "blank" to remove it).
References
Chalmers, T. C., & Lau, J. (1993). Meta-analytic stimulus for changes in clinical trials. Statistical Methods in Medical Research, 2(2), 161–172. https://doi.org/10.1177/096228029300200204
Lau, J., Schmid, C. H., & Chalmers, T. C. (1995). Cumulative meta-analysis of clinical trials builds evidence for exemplary medical care. Journal of Clinical Epidemiology, 48(1), 45–57. https://doi.org/10.1016/0895-4356(94)00106-z
Lewis, S., & Clarke, M. (2001). Forest plots: Trying to see the wood and the trees. British Medical Journal, 322(7300), 1479–1480. https://doi.org/10.1136/bmj.322.7300.1479
Viechtbauer, W. (2010). Conducting meta-analyses in R with the metafor package. Journal of Statistical Software, 36(3), 1–48. https://doi.org/10.18637/jss.v036.i03
See also
Examples
### calculate log risk ratios and corresponding sampling variances
dat <- escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg,
data=dat.bcg, slab=paste(author, year, sep=", "))
### fit random-effects model
res <- rma(yi, vi, data=dat)
### draw cumulative forest plots
x <- cumul(res, order=year)
forest(x)
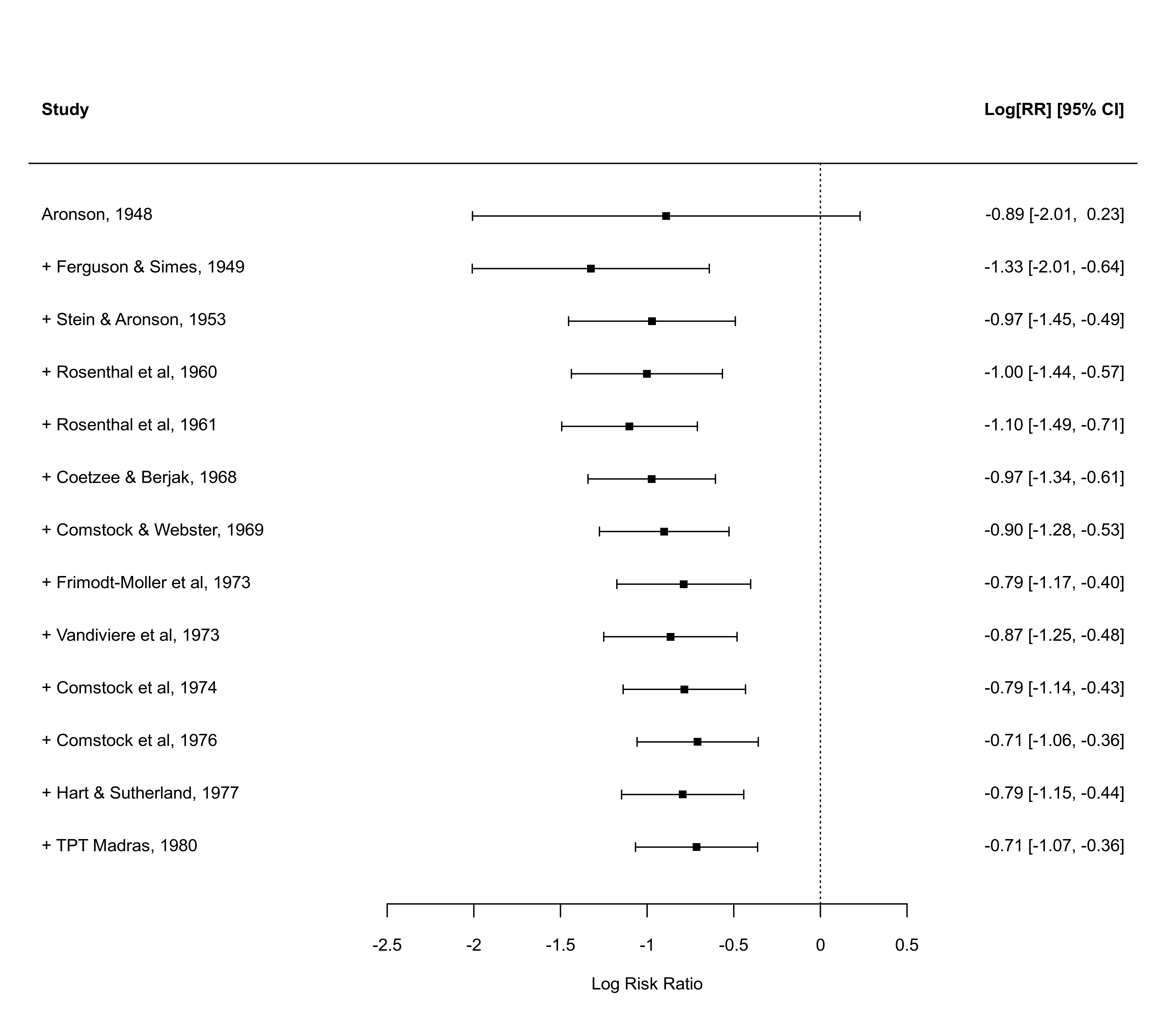 forest(x, xlim=c(-4,2.5), alim=c(-2,1), steps=7)
forest(x, xlim=c(-4,2.5), alim=c(-2,1), steps=7)
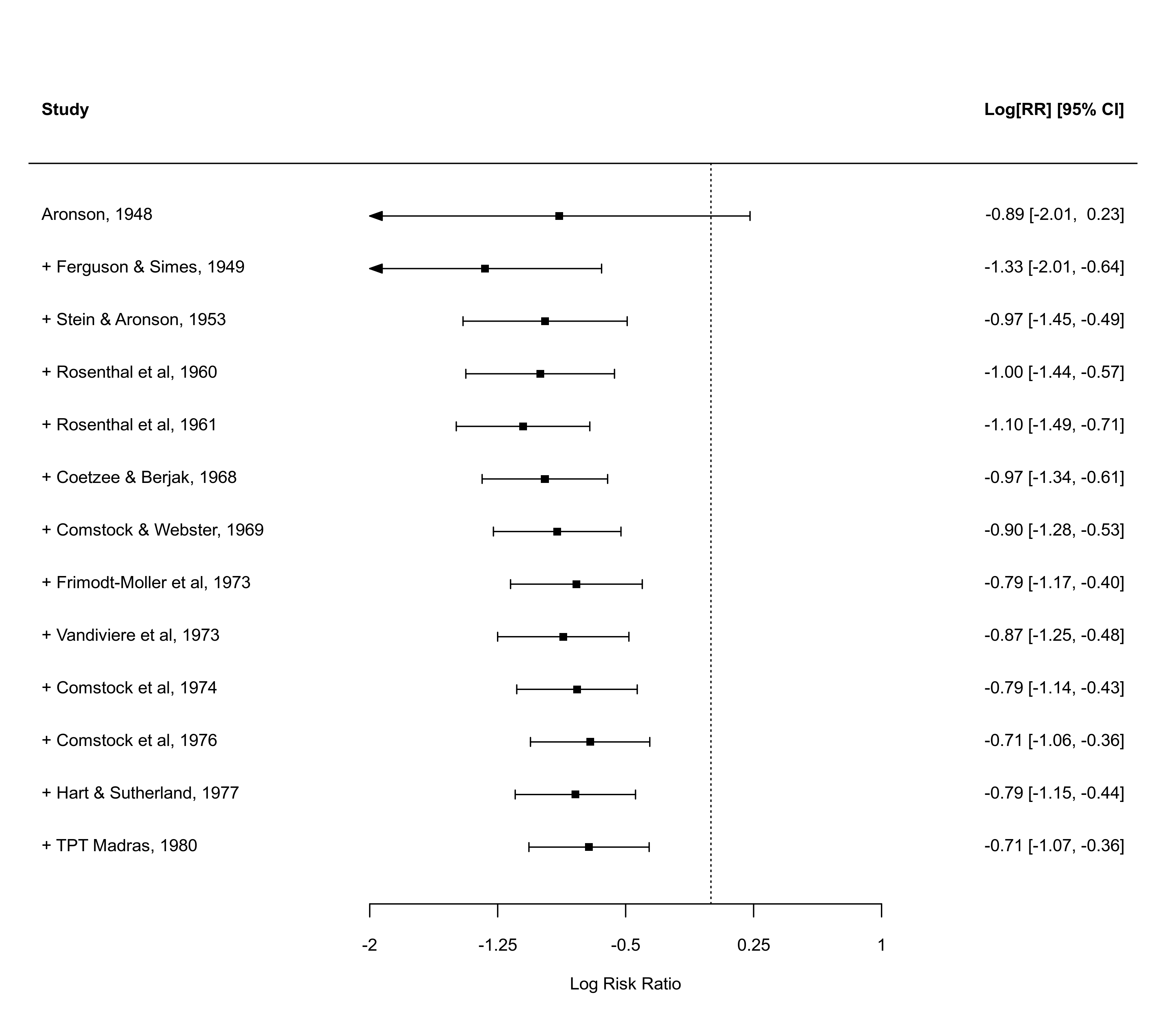 ### meta-analysis of the (log) risk ratios using the Mantel-Haenszel method
res <- rma.mh(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg,
slab=paste(author, year, sep=", "))
### draw cumulative forest plot
x <- cumul(res, order=year)
forest(x, xlim=c(-4,2.5), alim=c(-2,1), steps=7)
### meta-analysis of the (log) risk ratios using the Mantel-Haenszel method
res <- rma.mh(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg,
slab=paste(author, year, sep=", "))
### draw cumulative forest plot
x <- cumul(res, order=year)
forest(x, xlim=c(-4,2.5), alim=c(-2,1), steps=7)