Add Polygons to Forest Plots (Default Method)
addpoly.default.RdFunction to add one or more polygons to a forest plot.
# Default S3 method
addpoly(x, vi, sei, ci.lb, ci.ub, pi.lb, pi.ub,
rows=-1, level, annotate, predstyle, predlim, digits, width, mlab,
transf, atransf, targs, efac, col, border, lty, fonts, cex,
constarea=FALSE, ...)Arguments
- x
vector with the values at which the polygons should be drawn.
- vi
vector with the corresponding variances.
- sei
vector with the corresponding standard errors (note: only one of the two,
viorsei, needs to be specified).- ci.lb
vector with the corresponding lower confidence interval bounds. Not needed if
viorseiis specified. See ‘Details’.- ci.ub
vector with the corresponding upper confidence interval bounds. Not needed if
viorseiis specified. See ‘Details’.- pi.lb
optional vector with the corresponding lower prediction interval bounds.
- pi.ub
optional vector with the corresponding upper prediction interval bounds.
- rows
vector to specify the rows (or more generally, the positions) for plotting the polygons (defaults is
-1). Can also be a single value to specify the row of the first polygon (the remaining polygons are then plotted below this starting row). Whenpredstyleis not"line", can also be a vector of two numbers, the first for the position of the polygon, the second for the position of the prediction interval/distribution.- level
optional numeric value between 0 and 100 to specify the confidence interval level (see here for details).
- annotate
optional logical to specify whether annotations should be added to the plot for the polygons that are drawn.
- predstyle
character string to specify the style of the prediction interval (either
"line"(the default),"polygon","bar","shade", or"dist"; the last three only when adding a single polygon). Can be abbreviated.- predlim
optional argument to specify the limits of the predictive distribution when
predstyle="dist".- digits
optional integer to specify the number of decimal places to which the annotations should be rounded.
- width
optional integer to manually adjust the width of the columns for the annotations.
- mlab
optional character vector of the same length as
xgiving labels for the polygons that are drawn.- transf
optional argument to specify a function to transform the
xvalues and confidence interval bounds (e.g.,transf=exp; see also transf).- atransf
optional argument to specify a function to transform the annotations (e.g.,
atransf=exp; see also transf).- targs
optional arguments needed by the function specified via
transforatransf.- efac
optional vertical expansion factor for the polygons.
- col
optional character string to specify the color of the polygons.
- border
optional character string to specify the border color of the polygons.
- lty
optional argument to specify the line type for the prediction interval.
- fonts
optional character string to specify the font for the labels and annotations.
- cex
optional symbol expansion factor.
- constarea
logical to specify whether the height of the polygons (when adding multiple) should be adjusted so that the area of the polygons is constant (the default is
FALSE).- ...
other arguments.
Details
The function can be used to add one or more polygons to an existing forest plot created with the forest function. For example, pooled estimates based on a model involving moderators can be added to the plot this way (see ‘Examples’).
To use the function, one should specify the values at which the polygons should be drawn (via the x argument) together with the corresponding variances (via the vi argument) or with the corresponding standard errors (via the sei argument). Alternatively, one can specify the values at which the polygons should be drawn together with the corresponding confidence interval bounds (via the ci.lb and ci.ub arguments). Optionally, one can also specify the bounds of the corresponding prediction interval bounds via the pi.lb and pi.ub arguments. If the latter are specified, then they are added by default as lines around the summary polygons. When adding a single polygon to the plot, one can also use the predstyle argument to change the way the prediction interval is visualized (see forest.rma for details).
If unspecified, arguments level, annotate, digits, width, transf, atransf, targs, efac, fonts, cex, annosym, and textpos are automatically set equal to the same values that were used when creating the forest plot.
References
Viechtbauer, W. (2010). Conducting meta-analyses in R with the metafor package. Journal of Statistical Software, 36(3), 1–48. https://doi.org/10.18637/jss.v036.i03
See also
forest for functions to draw forest plots to which polygons can be added.
Examples
### calculate log risk ratios and corresponding sampling variances
dat <- escalc(measure="RR", ai=tpos, bi=tneg, ci=cpos, di=cneg, data=dat.bcg)
### fit mixed-effects model with absolute latitude as a moderator
res <- rma(yi, vi, mods = ~ ablat, slab=paste(author, year, sep=", "), data=dat)
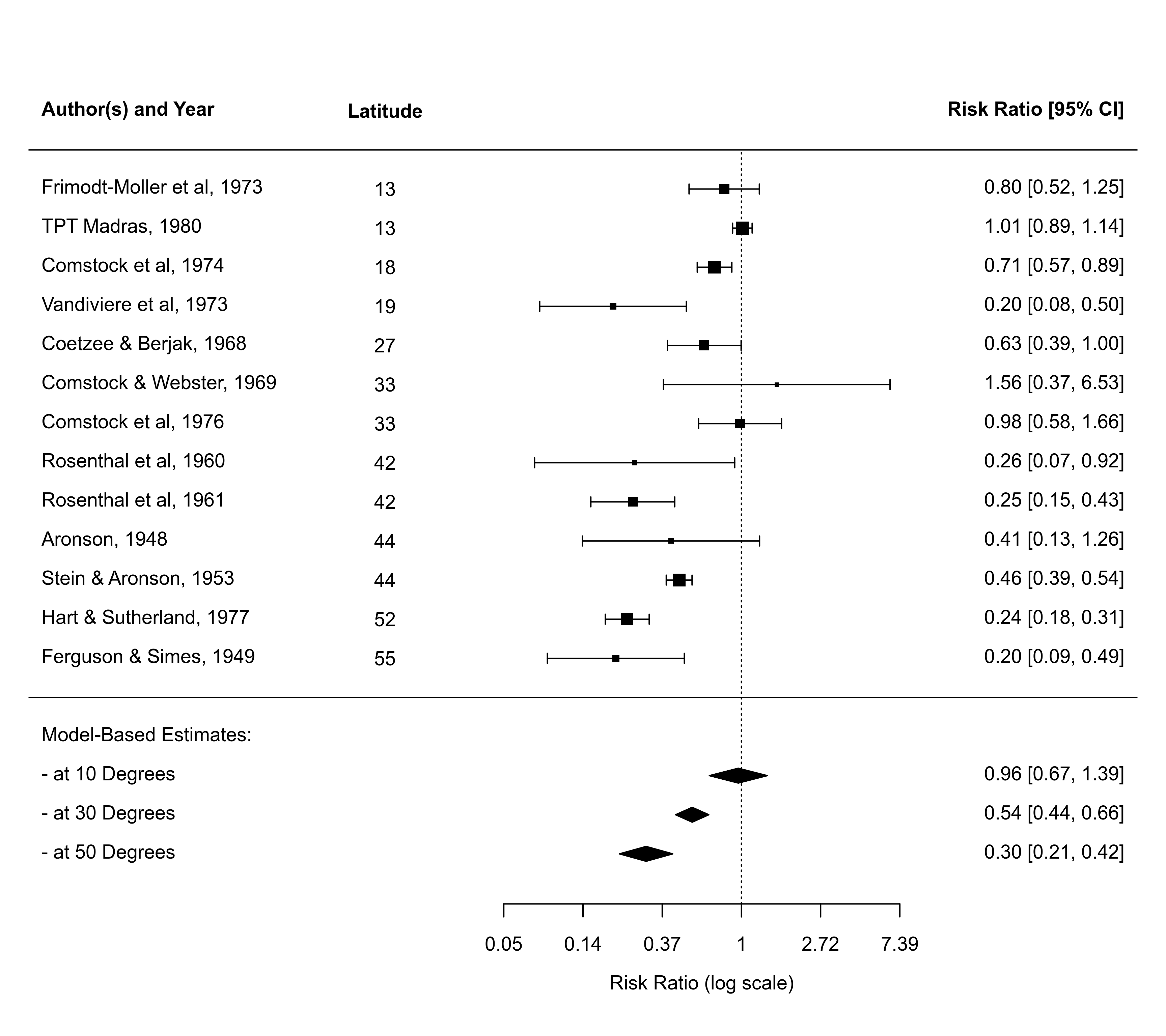
### forest plot of the observed risk ratios
forest(res, addfit=FALSE, atransf=exp, xlim=c(-9,5), ylim=c(-5,16), cex=0.9,
order=ablat, ilab=ablat, ilab.lab="Lattitude", ilab.xpos=-4.5,
header="Author(s) and Year")
### predicted average log risk ratios for 10, 30, and 50 degrees absolute latitude
x <- predict(res, newmods=c(10, 30, 50))
### add predicted average risk ratios to the forest plot
addpoly(x$pred, sei=x$se, rows=-2, mlab=c("- at 10 Degrees", "- at 30 Degrees", "- at 50 Degrees"))
abline(h=0)
text(-9, -1, "Model-Based Estimates:", pos=4, cex=0.9, font=2)