Studies on Intravenous Streptokinase for Acute Myocardial Infarction
dat.lau1992.RdResults from 33 trials comparing intravenous streptokinase versus placebo or no therapy in patients who had been hospitalized for acute myocardial infarction.
dat.lau1992Format
The data frame contains the following columns:
| trial | character | trial name |
| year | numeric | publication year |
| ai | numeric | number of deaths in the streptokinase group |
| n1i | numeric | number of patients in the streptokinase group |
| ci | numeric | number of deaths in the control group |
| n2i | numeric | number of patients in the control group |
Details
In the paper by Lau et al. (1992), the data are used to illustrate the idea of a cumulative meta-analysis, where the results are updated as each trial is added to the dataset. See ‘Examples’ for code that replicates the results and shows corresponding forest plots.
Source
Lau, J., Antman, E. M., Jimenez-Silva, J., Kupelnick, B., Mosteller, F., & Chalmers, T. C. (1992). Cumulative meta-analysis of therapeutic trials for myocardial infarction. New England Journal of Medicine, 327(4), 248–254. https://doi.org/10.1056/NEJM199207233270406
Concepts
medicine, cardiology, odds ratios, cumulative meta-analysis
Examples
### copy data into 'dat' and examine data
dat <- dat.lau1992
dat
#> trial year ai n1i ci n2i
#> 1 Fletcher 1959 1 12 4 11
#> 2 Dewar 1963 4 21 7 21
#> 3 European 1 1969 20 83 15 84
#> 4 European 2 1971 69 373 94 357
#> 5 Heikinheimo 1971 22 219 17 207
#> 6 Italian 1971 19 164 18 157
#> 7 Australian 1 1973 26 264 32 253
#> 8 Frankfurt 2 1973 13 102 29 104
#> 9 NHLBI SMIT 1974 7 53 3 54
#> 10 Frank 1975 6 55 6 53
#> 11 Valere 1975 11 49 9 42
#> 12 Klein 1976 4 14 1 9
#> 13 UK Collab 1976 38 302 40 293
#> 14 Austrian 1977 37 352 65 376
#> 15 Australian 2 1977 25 123 31 107
#> 16 Lasierra 1977 1 13 3 11
#> 17 N Ger Collab 1977 63 249 51 234
#> 18 Witchitz 1977 5 32 5 26
#> 19 European 3 1979 18 156 30 159
#> 20 ISAM 1986 54 859 63 882
#> 21 GISSI-1 1986 628 5860 758 5852
#> 22 Olson 1986 1 28 2 24
#> 23 Baroffio 1986 0 29 6 30
#> 24 Schreiber 1986 1 19 3 19
#> 25 Cribier 1986 1 21 1 23
#> 26 Sainsous 1986 3 49 6 49
#> 27 Durand 1987 3 35 4 29
#> 28 White 1987 2 107 12 112
#> 29 Bassand 1987 4 52 7 55
#> 30 Vlay 1988 1 13 2 12
#> 31 Kennedy 1988 12 191 17 177
#> 32 ISIS-2 1988 791 8592 1029 8595
#> 33 Wisenberg 1988 2 41 5 25
### load metafor package
library(metafor)
### meta-analysis of log odds ratios using the MH method
res <- rma.mh(measure="OR", ai=ai, n1i=n1i, ci=ci, n2i=n2i, data=dat, slab=trial)
print(res, digits=2)
#>
#> Equal-Effects Model (k = 33)
#>
#> I^2 (total heterogeneity / total variability): 18.98%
#> H^2 (total variability / sampling variability): 1.23
#>
#> Test for Heterogeneity:
#> Q(df = 32) = 39.50, p-val = 0.17
#>
#> Model Results (log scale):
#>
#> estimate se zval pval ci.lb ci.ub
#> -0.27 0.03 -8.16 <.01 -0.33 -0.20
#>
#> Model Results (OR scale):
#>
#> estimate ci.lb ci.ub
#> 0.76 0.72 0.82
#>
#> Cochran-Mantel-Haenszel Test: CMH = 66.60, df = 1, p-val < 0.01
#> Tarone's Test for Heterogeneity: X^2 = 43.84, df = 32, p-val = 0.08
#>
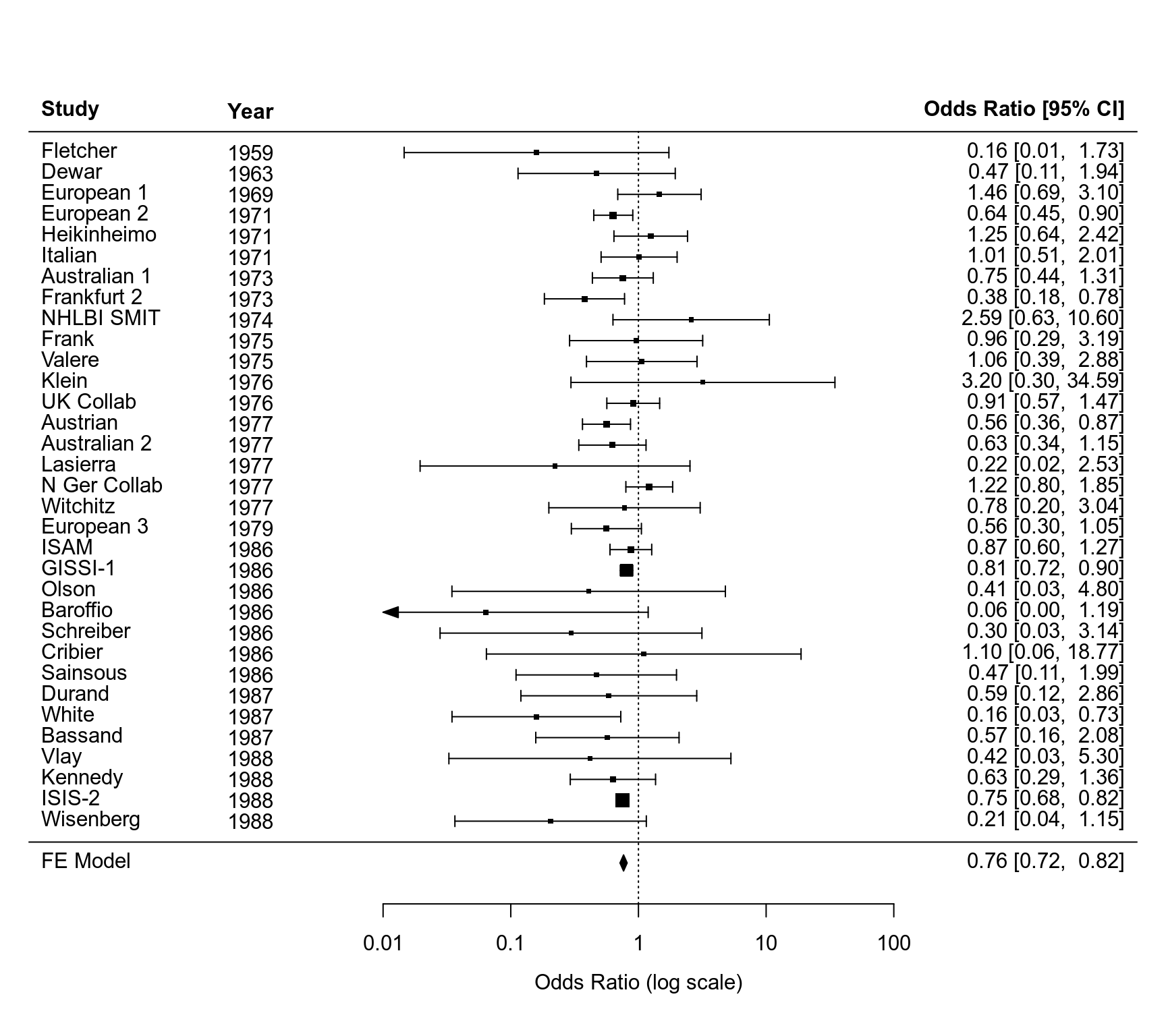
### forest plot
forest(res, xlim=c(-11,9), atransf=exp, at=log(c(0.01, 0.1, 1, 10, 100)),
ilab=dat$year, ilab.xpos=-7)
text(-7, 35, "Year", font=2)
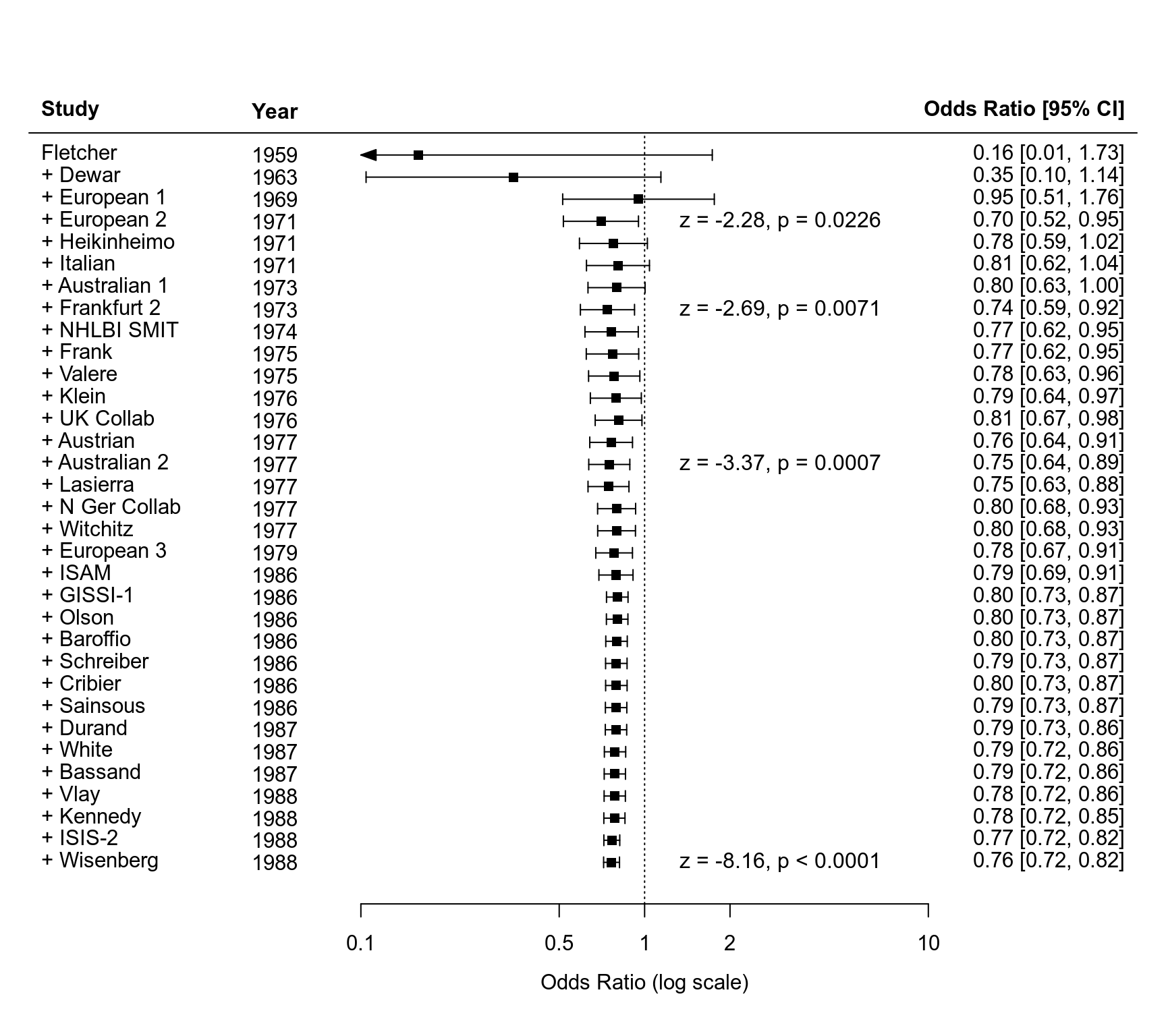
 ### cumulative meta-analysis
sav <- cumul(res)
### forest plot of the cumulative results
forest(sav, xlim=c(-5,4), atransf=exp, at=log(c(0.1, 0.5, 1, 2, 10)),
ilab=dat$year, ilab.xpos=-3)
text(-3, 35, "Year", font=2)
id <- c(4, 8, 15, 33) # rows for which the z/p-values should be shown (as in Lau et al., 1992)
text(1.1, (res$k:1)[id], paste0("z = ", fmtx(sav$zval[id], digits=2),
fmtp(sav$pval[id], pname=", p", equal=TRUE, sep=TRUE, add0=TRUE)))
### cumulative meta-analysis
sav <- cumul(res)
### forest plot of the cumulative results
forest(sav, xlim=c(-5,4), atransf=exp, at=log(c(0.1, 0.5, 1, 2, 10)),
ilab=dat$year, ilab.xpos=-3)
text(-3, 35, "Year", font=2)
id <- c(4, 8, 15, 33) # rows for which the z/p-values should be shown (as in Lau et al., 1992)
text(1.1, (res$k:1)[id], paste0("z = ", fmtx(sav$zval[id], digits=2),
fmtp(sav$pval[id], pname=", p", equal=TRUE, sep=TRUE, add0=TRUE)))